
Which would align with the assumption that colonisation precedes infection for many common bacterial infections, and that AMR prevalence in non-sterile site isolates mimics that in bloodstream infections www.nature.com/articles/s41...

Which would align with the assumption that colonisation precedes infection for many common bacterial infections, and that AMR prevalence in non-sterile site isolates mimics that in bloodstream infections www.nature.com/articles/s41...


Previous work led by Olga Tosas Auguet suggests yes: doi.org/10.1016/j.ec....
Hoping to innovate community population-level AMR surveillance with the ALARUM team bmjopen.bmj.com/content/16/2.... Can we use metagenomics of pooled colonisation samples to predict clinical AMR burden & inform antimicrobial guidelines?
Are you a post-doctoral microbiologist interested in characterising bacterial resistance evolution and want to apply your knowledge and expertise to developing new therapeutics for Klebsiella (and other Enterobacterales) infections? Then we need you! Apply now bit.ly/3MX39Ce close date March 13th 👀
🚨 Fully‑funded PhD for Home Students🚨
We'll explore non-antibiotic alternatives in relatistic environments, uncovering how they work, if they affect antibiotic activity and whether they influcence AMR acquistion. Come join a wonderful supervisory team!
#MicroSky #UTISky #AMar
tinyurl.com/38u552bu
🚨 New pre-print! 🚨 In the largest study of its kind to-date, we investigate the ecological and evolutionary mechanisms driving within-patient evolution of antimicrobial resistance (AMR). Read here:
www.biorxiv.org/content/10.6... , and follow along with this thread, discussing our findings (1/21)
If you're interested, click on www.ox.ac.uk/admissions/g... - Life and Medical Sciences, Clinical Medicine 04!

By Shireen Kotay/Amy Mathers - Biofilm removal in hospital sink drains drives unintended surges in antibiotic resistance rdcu.be/e1uTu. Effects akin to gut microbiome dysbiosis/loss of colonisation resistance. To mitigate these reservoirs we need better evidence to understand what works best where.


Our project "Sardinia’s Invisible Marine Microbiome" is one of six 2026 #PartArt4OW participatory art initiatives! Excited to do this with the brilliant @annadumitriu.bsky.social (image credit), Sardinian scientists, citizens of Alghero, and colleagues in @modmedmicro.bsky.social @bashthebug.net

Delighted to be involved in the Gram-Negative Antibiotic Discovery Innovator (Gr-ADI) consortium novonordiskfonden.dk/en/news/new-... as part of a project led by Annette von Delft gcgh.grandchallenges.org/grant/discov... @modmedmicro.bsky.social @samlipworth.bsky.social @philipwfowler.bsky.social

Univ of Oxford UNIQ+ scheme applications open: 7-week research internship this Summer www.ox.ac.uk/admissions/g.... Lots of projects - one in our lab on microplastics & AMR & hospital environments at @modmedmicro.bsky.social led by Aram Swinkels www.ox.ac.uk/admissions/g... (Clin Med 04). Join us!


New out by @dotnagy.bsky.social & colleagues synthesising published epidemiological/microbiological/genomic data on 'big 5' CPE hospital outbreaks: www.medrxiv.org/content/10.6...
Lots of them, but sampling and reporting heterogeneity makes it hard to nail risk factors for dissemination.

Our team wore blue for World Antimicrobial Awareness Week 2025💙
Research into how resistance emerges, spreads and can be prevented is vital. Our group works to understand these processes and develop better tools to detect, monitor and combat antimicrobial resistance.
#WAAW @oxfordbrc.bsky.social
Congrats Viki!

Pleased to see this out today in @lancetmicrobe.bsky.social www.sciencedirect.com/science/arti.... We estimate the independent effects of AMR genes on MIC at a drug-bug level and challenge the idea that resistance is a binary phenomenon.
Now published in @natcomms.nature.com 🎉
www.nature.com/articles/s41...
With Gillian Rodger, @nstoesser.bsky.social, @samlipworth.bsky.social, @stat-sarah.bsky.social, and many others!
Pleased to see this pre-printed, highlighting the completeness/accuracy of @nanoporetech.com long-read genome assembly for clinical Enterobacterales: www.biorxiv.org/content/10.1...
Thanks to colleagues @modmedmicro.bsky.social, @ukhsa.bsky.social, @genewiz.bsky.social and @oxfordbrc.bsky.social!
Job opportunity for microbial genomicists interested in E. coli and Klebsiella infections and AMR 👀⬇️ - do get in touch if you might be interested! Closing date 7th Oct.
@modmedmicro.bsky.social
Great new opportunity to work with #Liamshaw and @mutantbug.bsky.social as part of our HPRU collaboration (www.expmedndm.ox.ac.uk/research/hpr...) - sinks, phages, wastewater genomes, novel threat detection bit.ly/45Wcv6I - deadline 14th Sept 👀
More evidence to support best practice "water-safe" care in healthcare facilities is urgently needed. If you're interested please tap into the @hisinfection.bsky.social Wastewater and AMR Special Interest Collaborative (SPARC) - see you there!

More discussion about the JHI study with @emrsa15.bsky.social in the Infection Control Matters podcast series infectioncontrolmatters.com/2025/04/02/t...
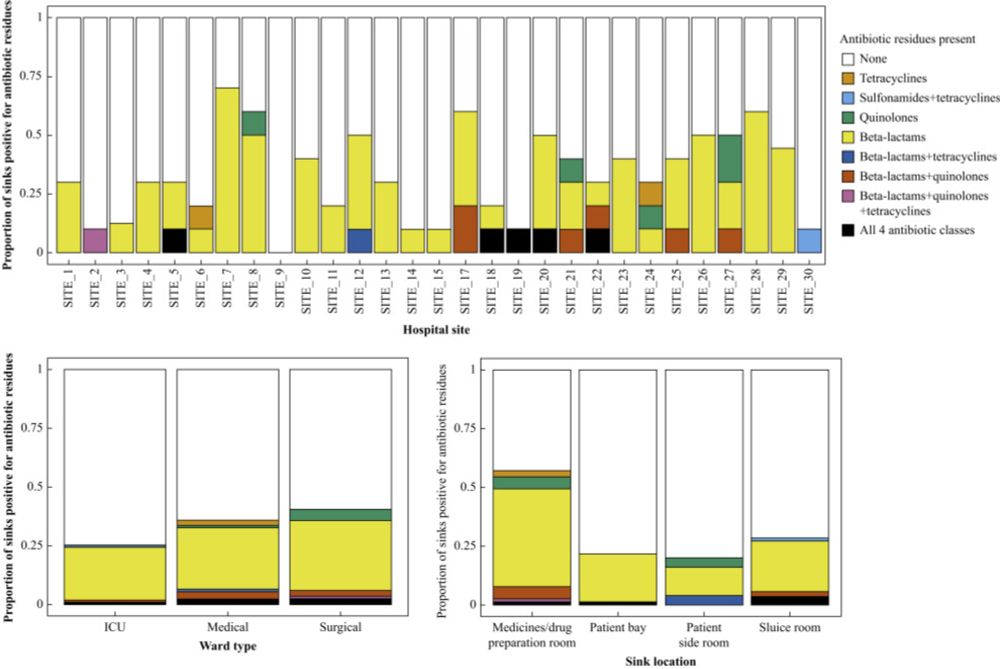
This builds on the SinkBug group's earlier work in @thejhi.bsky.social www.journalofhospitalinfection.com/article/S019... where we showed that sink infrastructure in hospitals was common, highly varied, and that antibiotics were detected in a third of sink-traps.
New analysis from Kevin Chau @modmedmicro.bsky.social for the SinkBug Consortium (a collaboration with teams across 29 UK hospitals, NITCAR, UKHSA): Pathogen and AMR gene burden in 287 hospital sink-trap metagenomes and factors associated with high burdens.