Interested in transcriptional regulation, enhancers and 3D genome folding?
In this new study we wondered about the role of cohesin loading at enhancers for long-range transcriptional control
www.biorxiv.org/content/10.6...
detailed 🧵👇
12.02.2026 21:39
👍 67
🔁 33
💬 1
📌 3
Happy to share part of my postdoctoral work at the @lucas.farnunglab.com lab. Great collaboration with @voslab.org and @andersshansen.bsky.social. “Structural basis for CTCF-mediated chromatin organization” www.biorxiv.org/content/10.6...
09.02.2026 15:22
👍 30
🔁 10
💬 0
📌 0
Very excited to share our new paper 🎊
A wonderful early birthday gift! 🎂🎁
Huge thanks to everyone who contributed to making this possible.
16.12.2025 09:23
👍 5
🔁 1
💬 0
📌 0
Here is a copy of last year's Twitter thread explaining our preprint - jump to (21) for the new stuff 👀
Synergy between cis-regulatory elements can render cohesin dispensable for distal enhancer function
now revised and journal accepted at www.science.org/doi/10.1126/...
🧵👇
27.11.2025 21:58
👍 89
🔁 45
💬 4
📌 3

📣 Paper alert!
I am delighted that our paper exploring the impact of Neanderthal-derived variants on the activity of a disease-associated craniofacial enhancer has been published in Development today!
journals.biologists.com/dev/article/...
10.11.2025 11:11
👍 123
🔁 49
💬 7
📌 6

How can we organize current theoretical approaches for developmental biology - from information to dynamical systems & GRNs - into a common framework?
We propose to think along Marr's 3 levels: computational problem, algorithm, implementation
Check out our review:
arxiv.org/abs/2510.24536
10.11.2025 06:46
👍 49
🔁 13
💬 1
📌 2

🥁This Wednesday , in #FragileNucleosome seminar, we are excited to host @hannahlong.bsky.social and @jeffvierstra.bsky.social to tell us about amazing work they are doing!
🗓️Register here for upcoming session and the entire series:
us06web.zoom.us/webinar/regi...
03.11.2025 12:03
👍 20
🔁 12
💬 0
📌 0

Characterization of induced cohesin loop extrusion trajectories in living cells
Nature Genetics - This study introduces a system called TArgeted Cohesin Loader (TACL) that recruits cohesin complexes at defined genomic regions and induces loop extrusion events in living cells,...
The TArgeted Cohesin Loader (TACL) paper was just published. Happy that we were able to contribute to this really exciting project!
If you want to learn how targeting cohesin to defined loci in the genome affects the local chromatin environment and transcription, look no further!
rdcu.be/eLiT5
16.10.2025 20:17
👍 54
🔁 20
💬 0
📌 0

Happy to share our latest publication, in which we show that the arrangement of nucleosomes around CTCF sites contributes to higher-order organisation of chromatin into TADs: www.embopress.org/doi/full/10....
27.10.2025 11:49
👍 71
🔁 31
💬 0
📌 2
Out now! 🎉 Check the thread & preprint to see why we think E–P specificity is real in mammals — and, well, a few other interesting things popped up too 👀
Huge thanks to @danielibrahim.bsky.social, @arnaudkr.bsky.social & @stemundi.bsky.social and fantastic people in their labs — what a journey! 🧪🔬
17.10.2025 05:50
👍 43
🔁 13
💬 1
📌 0
Really excited to share our latest work led by @mattiaubertini.bsky.social and @nesslfy.bsky.social: we report that cohesin loop extrusion creates rare but long-lived encounters between genomic sequences which underlie efficient enhancer-promoter communication.
www.biorxiv.org/content/10.1...
A🧵👇
24.09.2025 21:45
👍 114
🔁 52
💬 8
📌 5
Have you ever wondered how the exact location of a gene affects it's activity?
The main story of my PhD deals with exactly that question, and is now published in Science! ✨
www.science.org/doi/10.1126/...
My amazing co-author and friend @mathiaseder.bsky.social summarized the highlights for you
19.09.2025 14:22
👍 36
🔁 13
💬 1
📌 2
Very proud of our paper on "scrambling-by-hopping" LADs, which was just published: www.nature.com/articles/s41.... Congrats to Lise Dauban and the rest of the team – this was a real tour-de-force!
02.09.2025 17:26
👍 58
🔁 24
💬 0
📌 0
This is not a HiC map! Ever wondered if multiple enhancers get activated simultaneously? We measured chromatin accessibility on thousands of molecules by nanopore to create genome-wide co-accessibility maps. Proud of @mathias-boulanger.bsky.social @kasitc.bsky.social Biology in the thread👇
18.08.2025 13:00
👍 113
🔁 34
💬 10
📌 0
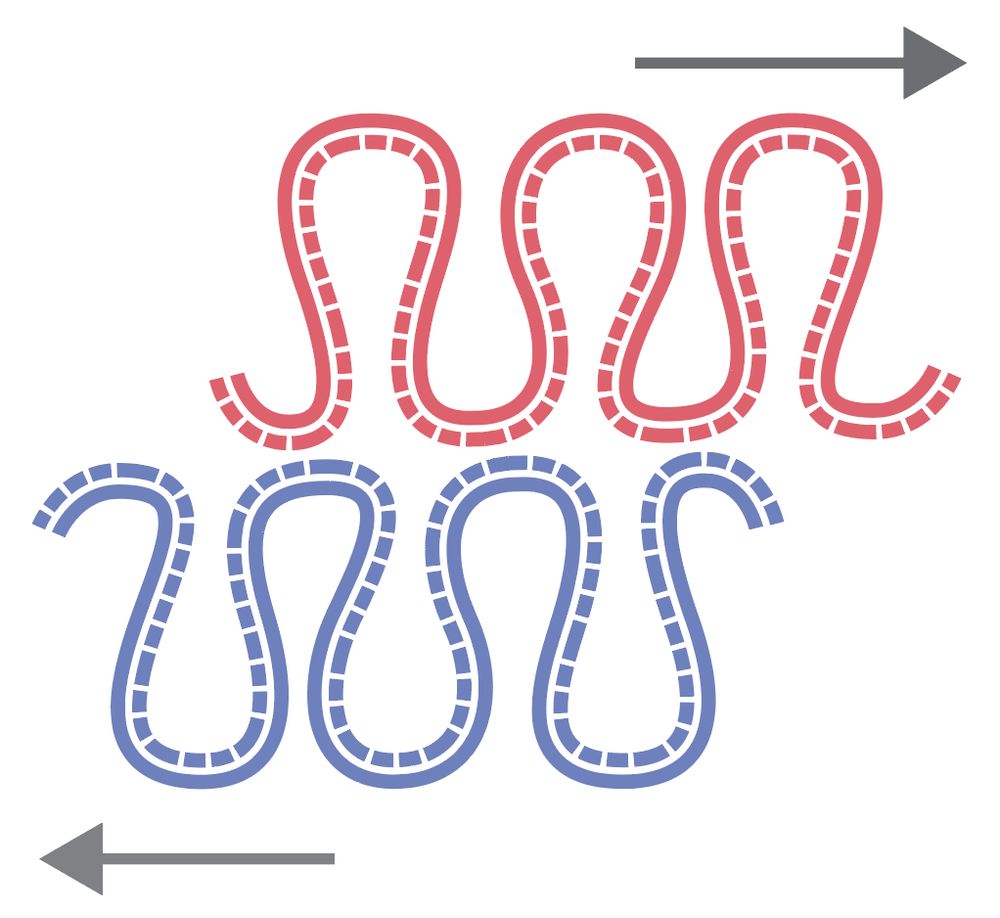
We found a new asymmetry in the large-scale chromosome structure: sister chromatids are systematically shifted by hundreds of kb in the 5′→3′ direction of their inherited strands! The work was led by Flavia Corsi, in close collaboration with the Daniel Gerlich lab.
www.biorxiv.org/content/10.1...
1/
15.07.2025 08:11
👍 115
🔁 59
💬 3
📌 7
Calling all aspiring Postdocs! One extra week to apply to join our exciting HFSP-funded project to uncover how chromatin moves to function.
Deadline July 15th.
www.mdc-berlin.de/career/jobs/...
08.07.2025 09:12
👍 15
🔁 16
💬 0
📌 0
Friedrich Miescher Institute, open position
Friedrich Miescher Institute, open position
We are looking for a research assistant to join our lab and support ongoing projects with genome engineering in mouse embryonic stem cells! Please get in touch if you are interested.
www.fmi.ch/education-ca...
26.06.2025 15:04
👍 19
🔁 22
💬 0
📌 0
Very happy to share the peer-reviewed version of our paper in which we study the formation and function of pair-wise and multi-way enhancer-promoter interactions in gene regulation (see thread below): www.nature.com/articles/s41...
13.05.2025 09:41
👍 103
🔁 32
💬 3
📌 3
Rules of Engagement
YouTube video by Johan Gibcus
Earnshaw, Goloborodko, Dekker & Mirny labs are excited to present our latest work, "Rules of engagement for condensins and cohesins guide mitotic chromosome formation" - now accepted!!
www.science.org/doi/10.1126/...
A short clip describing the key results:
www.youtube.com/watch?v=pmvO...
11.04.2025 07:45
👍 98
🔁 53
💬 7
📌 6

Happy to share the latest story from @arnaudkr.bsky.social's lab @embl.org! With @guidobarzaghi.bsky.social, we used Single Molecule Footprinting to quantify how often chromatin is accessible at enhancers after TF and chromatin environment changes! Check our preprint bit.ly/3XQMFxN + thread ⬇️ 1/11
08.04.2025 13:51
👍 78
🔁 33
💬 4
📌 2

The importance of moving away from bulk! Occupancy of Pol II at promoters is dramatically different between fly and mouse cells! When looking single molecule! Proud of the team! @kasitc.bsky.social @molinalab.bsky.social doi.org/10.1038/s443...
02.04.2025 08:47
👍 115
🔁 37
💬 2
📌 1

Celebrating 10 years of our lab with a new preprint:
www.biorxiv.org/content/10.1...
How does enhancer location within a TAD control transcriptional bursts from a cognate promoter?
Experiments by Jana Tünnermann and modelling by Gregory Roth
29.03.2025 12:45
👍 149
🔁 58
💬 6
📌 2


















