
I had a great time nerding out about protein design with @nickpolizzi.bsky.social, and an enormous thanks to @biotechtv.bsky.social and @massbio.bsky.social for hosting us!
www.biotechtv.com/post/chris-b...

I had a great time nerding out about protein design with @nickpolizzi.bsky.social, and an enormous thanks to @biotechtv.bsky.social and @massbio.bsky.social for hosting us!
www.biotechtv.com/post/chris-b...
6/6
CANVAS turns minimal TIM barrels into blank canvases for enzyme design. This approach could be applied to other minimal folds, enabling de novo enzymes for virtually any reaction while keeping stable, well-behaved scaffolds. The goal: bespoke enzymes for chemistries nature hasn’t explored yet!

5/6
Molecular dynamics simulations helped dissect structural determinants of catalysis. Our catalytic‑competency score, i.e. the fraction of snapshots with both key catalytic contacts formed, tracks well with experimental activity across variants.

4/6
To demonstrate evolvability, we applied our previously reported ensemble‑based design method (pubs.acs.org/doi/full/10....) to the low‑efficiency variant KempTIM4, boosting kcat/KM >1,600‑fold (to 32,000 M⁻¹ s⁻¹) with only 8 additional variants screened.

3/6
We solved the KempTIM1 crystal structure (blue) bound to a transition‑state analogue. The designed lid (magenta) shows excellent agreement with the model, and the ligand binds in a similar pose to the design model, enabling the intended catalytic contacts.

2/6
After reviewer feedback, we boosted activity and hit rate. Keeping the same active‑site design, we improved solubility and expression via surface redesign, yielding KempTIM1, one of the most active first‑round de novo enzymes reported to date (kcat/KM = 21,000 M⁻¹ s⁻¹).

1/6
Thrilled to announce our updated preprint on the CANVAS method for de novo enzyme design, in collaboration with @birtehoecker.bsky.social. We show how CANVAS builds custom lids onto minimal TIM barrels, transforming them into efficient catalysts.
www.biorxiv.org/content/10.1...

Announcement for the 6th Protein Engineering Canada Conference, to be held June 22nd-24th in Ottawa, Canada. The image shows a protein structure in front of a picture of Parliament Hill and Chateau Laurier in Ottawa.
1/
Save the date!
The 6th Protein Engineering Canada Conference will be held on June 22-24 in Ottawa, Canada.
Abstract submission and registration are open!
More information here: event.fourwaves.com/pec2026/pages

2/
Speakers of the 6th Protein Engineering Canada Conference include:
Bill DeGrado
Joelle Pelletier
Tim Whitehead
Anastassia Vorobieva
Lucy Colwell
Ai Niitsu
@joannas.bsky.social
@nickpolizzi.bsky.social
@possuhuanglab.bsky.social
@paolalaurino.bsky.social
@stephanhammer.bsky.social
and more!

Announcement for the 6th Protein Engineering Canada Conference, to be held June 22nd-24th in Ottawa, Canada. The image shows a protein structure in front of a picture of Parliament Hill and Chateau Laurier in Ottawa.
1/
Save the date!
The 6th Protein Engineering Canada Conference will be held on June 22-24 in Ottawa, Canada.
Abstract submission and registration are open!
More information here: event.fourwaves.com/pec2026/pages

Thrilled to share that our latest article is now out in final form! A great collaboration with @fraserlab.com and @silviaosuna.bsky.social.
Distal mutations enhance catalysis in designed enzymes by facilitating substrate binding and product release
www.nature.com/articles/s41...

Our latest article is now published online! In collaboration with @thompson-lab.bsky.social, Marc Garcia-Borràs, and @ferranfeixas.bsky.social.
Distal Mutations in a Designed Retro-Aldolase Alter Loop Dynamics to Shift and Accelerate the Rate-Limiting Step
pubs.acs.org/doi/full/10....

Similarly, enzyme function can be designed de novo by creating a new active site within a natural protein scaffold that lacks the target activity, even if that catalytic function exists in nature.
See below for an early example:
www.pnas.org/doi/full/10....
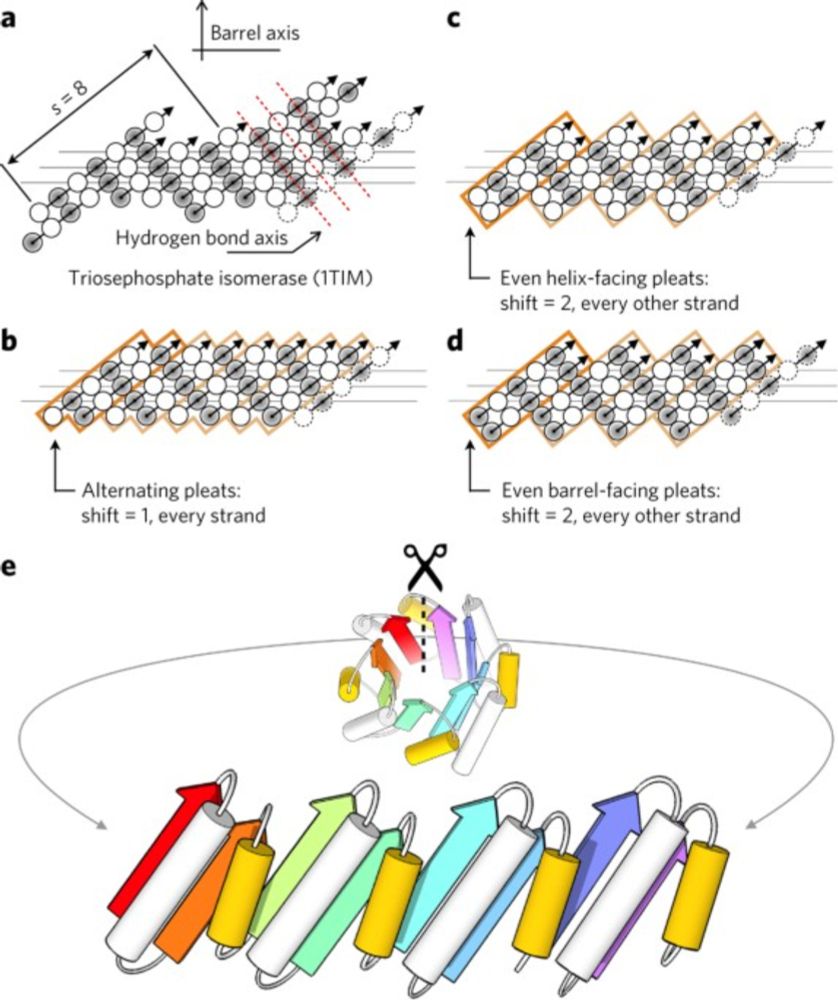
No, I don’t think that’s necessarily implied. For example, a TIM barrel can be designed from scratch without referencing any specific natural sequence or structure, even if this fold exists in nature. I consider this de novo design. See example below:
www.nature.com/articles/nch...

The design and creation of a protein sequence, structure or function from scratch, rather than modifying a pre-existing sequence, structure or function.
An early pioneer of this field is Bill DeGrado, see below.
www.science.org/doi/10.1126/...
Overall, our study:
✅ Introduces a new strategy to transform minimal protein scaffolds into biocatalysts
✅ Provides mechanistic insights from crystallography & molecular dynamics
✅ Opens the door to designing custom lids for more complex reactions, which we’re now exploring
Thanks for reading! 🧵🧬
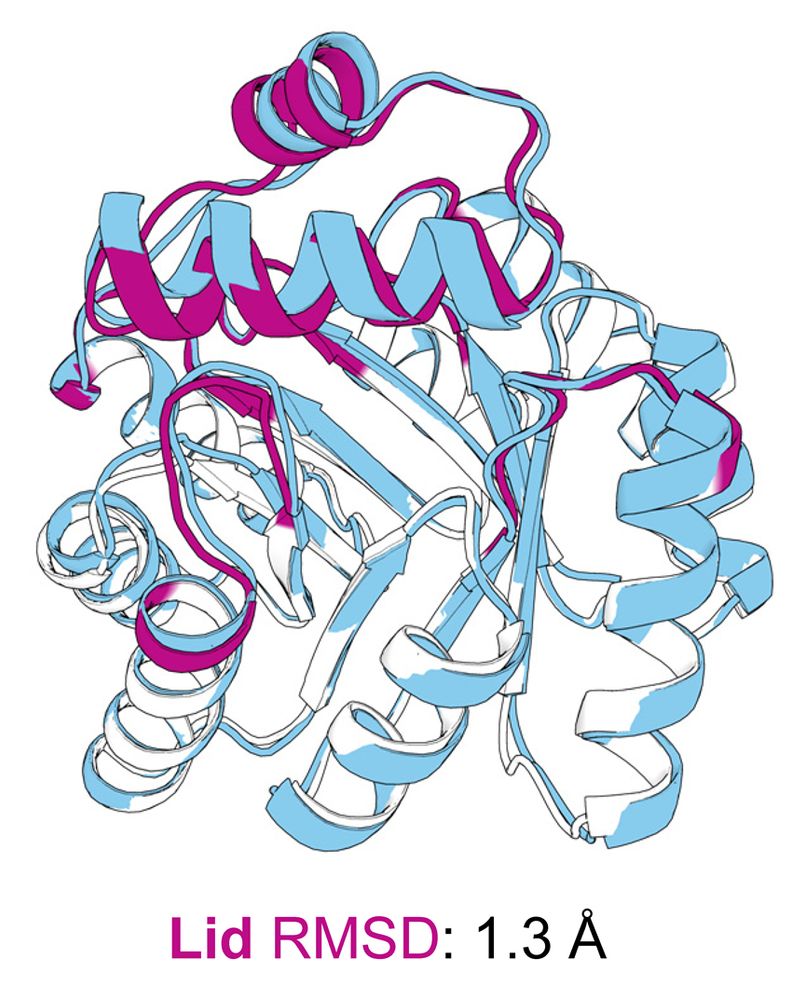
The crystal structure (blue) aligns closely with the design model (minimal TIM barrel and lid colored white and magenta, respectively).
Our crystal structure validated the designed fold, confirming that the lid was correctly folded.
However, a subtle 1.8 Å lid shift disrupted a key catalytic contact, likely contributing to the modest activity. But structural analysis reveals paths to improve activity in the next round of design!
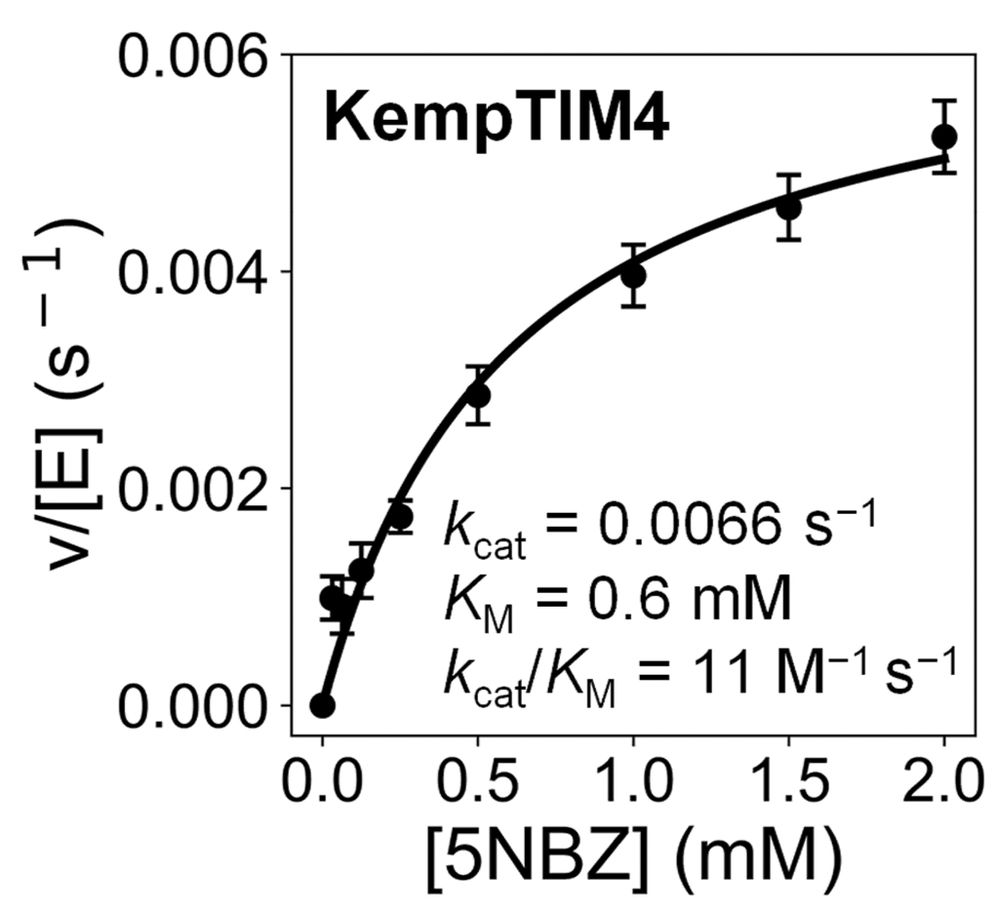
Michaelis-Menten plot of KempTIM4 showing saturation kinetics.
One of our designs, KempTIM4, showed catalytic efficiency comparable to many first-round de novo Kemp eliminases generated by traditional methods.
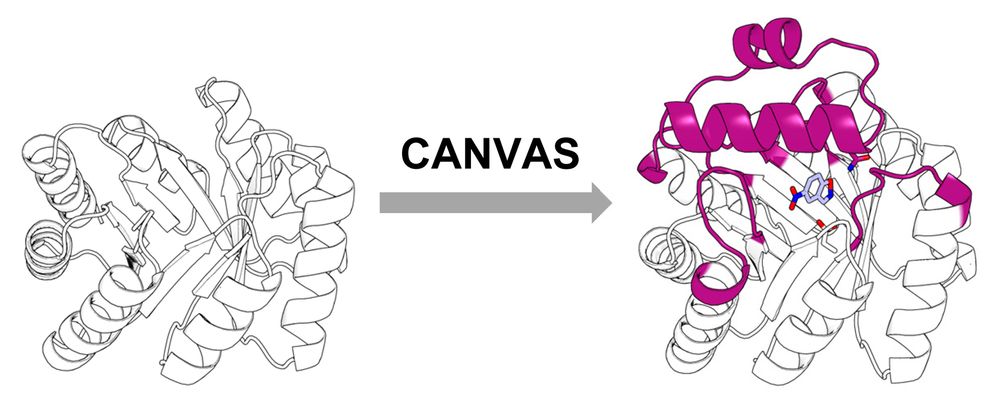
Building a custom lid onto a minimal, de novo TIM barrel using CANVAS.
Using CANVAS, we designed a structural lid onto a minimal, de novo TIM barrel to anchor catalytic residues and create an active site for the Kemp elimination reaction.
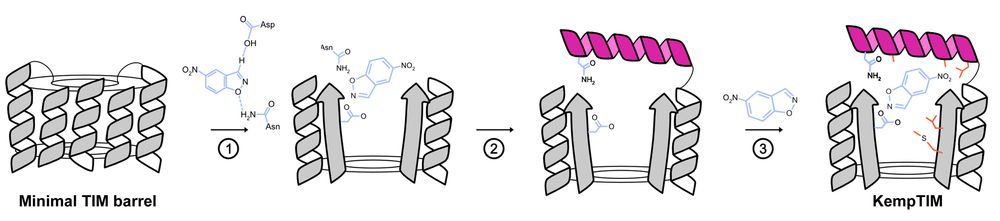
De novo enzyme design using CANVAS.
TIM barrels are among nature’s most powerful enzyme scaffolds but making them from scratch with catalytic function has been a challenge.
Enter CANVAS: a computational pipeline combining Triad, RFdiffusion & ProteinMPNN to customize minimal TIM barrels into functional enzymes.

In collaboration with @birtehoecker.bsky.social, we’ve unlocked enzymatic activity in a minimal de novo TIM barrel by designing a custom lid for catalysis! 🧵👇
#ProteinDesign #EnzymeDesign
Customizing the Structure of a Minimal TIM Barrel to Create a De Novo Enzyme
www.biorxiv.org/content/10.1...
Congratulations! Looking forward to seeing all the exciting science that will come out of your lab! 🧪

Join us! We are looking for a new team member (PhD student) with strong background in organic chemistry.
🙏 RETWEET (We want to recruit internationally)
Organic chemistry meets #DirectedEvolution
Highly interdisciplinary & passionate research group
uni-bielefeld.hr4you.org/job/view/433...

Non-canonical amino acid from PDB ID 8W3Z shown chelating to a magnesium ion. Image made with PyMol. https://www.rcsb.org/structure/8W3Z https://www.pymol.org
Protein Engineering, Design & Selection (PEDS) invites contributions to a Special Collection titled, “Non-Canonical Amino Acids", with guest editors Prof. Huiwang Ai (Virginia) and Prof. Peng Chen (Peking). Send us your best work!
academic.oup.com/peds/pages/c...

Super excited to share a new preprint from our lab on design of small-molecule binding proteins using neural networks! The paper has a bit of everything. A new graph neural network, new design algorithms, and experimental validation. www.biorxiv.org/content/10.1...
🧵🧪

Guess what? TPS has extended the deadline to March 19 to submit abstracts for poster presentations and speaking opportunities at our 39th Annual Symposium. Join us in San Francisco June 26 - 29 for 3.5 days of scientific talks.
hashtag#proteinscience hashtag#annualsymposium
lnkd.in/g7VKqX7C

The take-home message? Distal residues actively shape enzyme catalysis. Optimizing them can remove bottlenecks in substrate binding & product release—boosting activity. Want to dive deeper? Read our full study here: www.biorxiv.org/content/10.1...
(6/6)
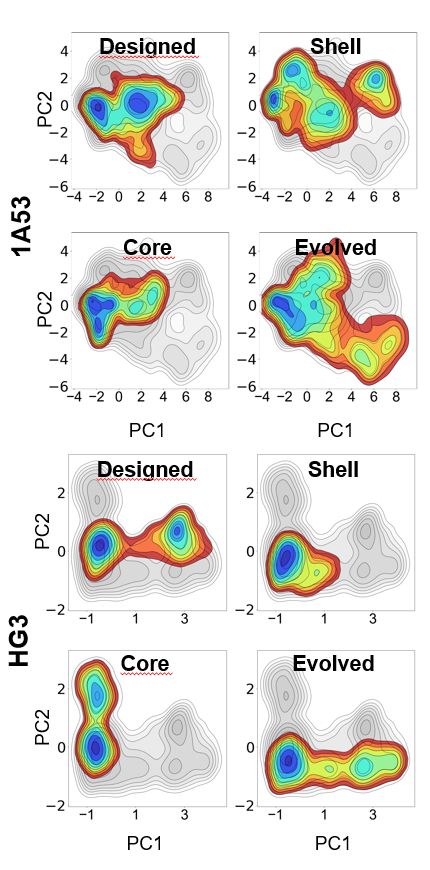
Molecular dynamics simulations showed that distal mutations enhance active-site accessibility—either by loosening loops covering the active site or widening bottlenecks for substrate entry & product exit. The enzyme breathes more efficiently! 🌬️ (5/6)

Kinetic solvent viscosity effects & stopped-flow experiments showed that distal mutations don’t just tweak structure—they accelerate substrate binding & product release. (4/6)
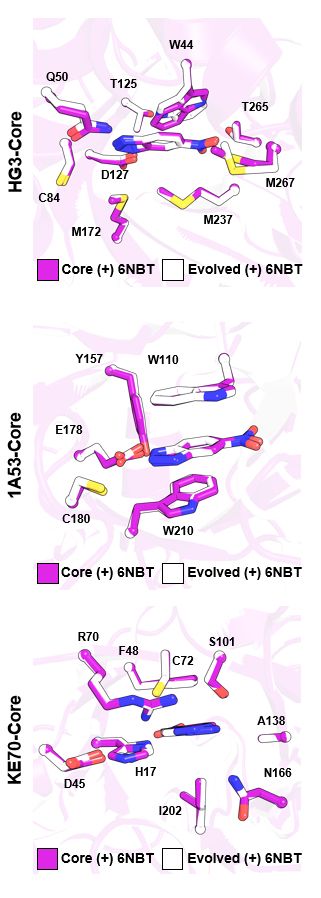
Crystal structures showed that active-site mutations pre-organize the catalytic machinery. But distal mutations? They subtly tune conformational dynamics—enhancing productive substates & reshaping the energy landscape of the catalytic cycle. (3/6)