Scientific American has updated the figure, now grouped into swimmers, fliers, walkers/runners, and vehicles. A person on a bicycle remains the most efficient way to travel, compared to all forms of biological locomotion and mechanical transport.
www.scientificamerican.com/article/a-hu...
04.03.2026 11:52
👍 568
🔁 210
💬 17
📌 31

Estimating Organism Abundance Using Within‐Sample Haplotype Frequencies of eDNA Data
Environmental DNA (eDNA) provides powerful insights into species presence and community composition but remains limited in its capacity to infer species abundance or population structure. Here, we sh...
🧬 My haplotype paper is out!
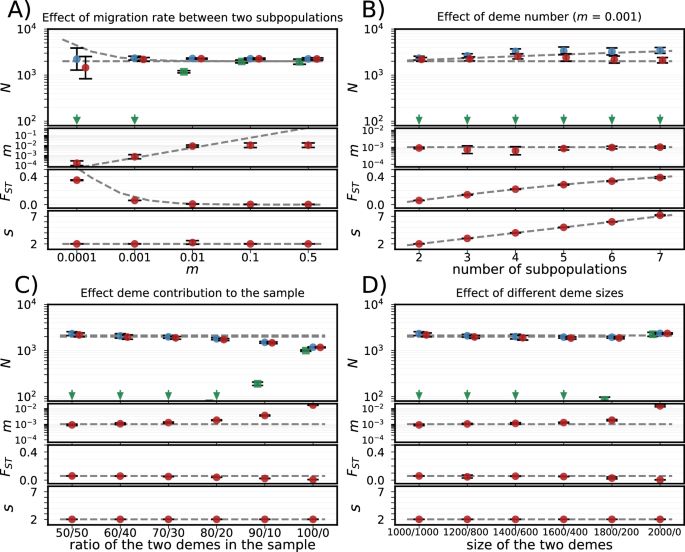
We show that deviations between within-sample and population-level haplotype frequencies can be used to estimate how many individuals contributed to an eDNA sample.
No tissue references needed, just metabarcoding data and some population genetics.
#eDNA #PopGen
13.02.2026 16:54
👍 42
🔁 15
💬 2
📌 1
Final version of paper with @smishra677.bsky.social now published in a wonderful issue of GENETICS!
academic.oup.com/genetics/art...
08.01.2026 13:57
👍 26
🔁 16
💬 0
📌 0

Not Just Ne Ne-More: New Applications for SMC from Ecology to Phylogenies
Abstract. Genomes contain the mutational footprint of an organism’s evolutionary history, shaped by diverse forces including ecological factors, selective
In a new GBE Review, @david-peede.bsky.social et al. overview the SMC model and extensions, discuss examples of discoveries made with the help of SMC-based inference, and comment on the assumptions, benefits, and drawbacks of various methods.
🔗 doi.org/10.1093/gbe/...
#genome #evolution #compbio
08.01.2026 08:05
👍 36
🔁 20
💬 1
📌 2

Finally, we’ve solved a long-standing mystery: what tintinnid shells are actually made of:
A new class of biomaterial formed by remarkable structural proteins unique to tintinnids.
A major milestone after 3 years of work! Read about it in our preprint: doi.org/10.64898/202...
#ProtistsOnSky
27.12.2025 10:30
👍 95
🔁 35
💬 4
📌 3

Genomic insights into the origin of ecotypes
A century after Turesson introduced ecotypes as locally adapted subunits of species, their origins and nature remain debated. Recent advances in molecular ecology have breathed new life into the ecotype concept and significantly expanded our understanding of how ecotypes form. Here, we clarify how ecotypes can be distinguished from other forms of local adaptation, outline criteria for their recognition, and synthesise evidence on the ecological and evolutionary processes that give rise to them. We then evaluate what genomics has uncovered about the origin of ecotypes, revealing pronounced roles of pre-existing variation, large-effect loci, structural variants, gene regulation, and complex demographic histories. Our framework refines Turesson’s concept and highlights outstanding questions about the predictability, persistence, and evolutionary significance of ecotypes.
Online now: Genomic insights into the origin of ecotypes
15.12.2025 12:56
👍 6
🔁 1
💬 0
📌 0

GHIST logo
Congratulations to the winners of the 2025 Genomic History Inference Strategies Tournament challenges!
Among the 9 challenges, we had five winners: @alwaysrong.bsky.social, @adaigle.bsky.social, @andrewhvaughn.bsky.social, @thymelicus.bsky.social, @rgollnisch.bsky.social
15.12.2025 18:48
👍 20
🔁 10
💬 2
📌 0


Obsessed with this Journal of Immaterial Science article 🔥
08.10.2025 08:56
👍 65
🔁 11
💬 2
📌 4
Low-coverage sequencing (LCS) + genotype imputation is a cost-effective approach to genotype hundreds of samples at a full-genome level. But how do different imputation tools perform across populations with varying relatedness and inbreeding?
04.10.2025 08:11
👍 10
🔁 6
💬 1
📌 0
Check out our new paper on adopting a trait-based framework for protist diversity! We make the case for a unified protist trait database, how to build it, and how it could transform research on protist ecology and evolution.
#protistsonsky
03.09.2025 08:47
👍 58
🔁 27
💬 2
📌 1
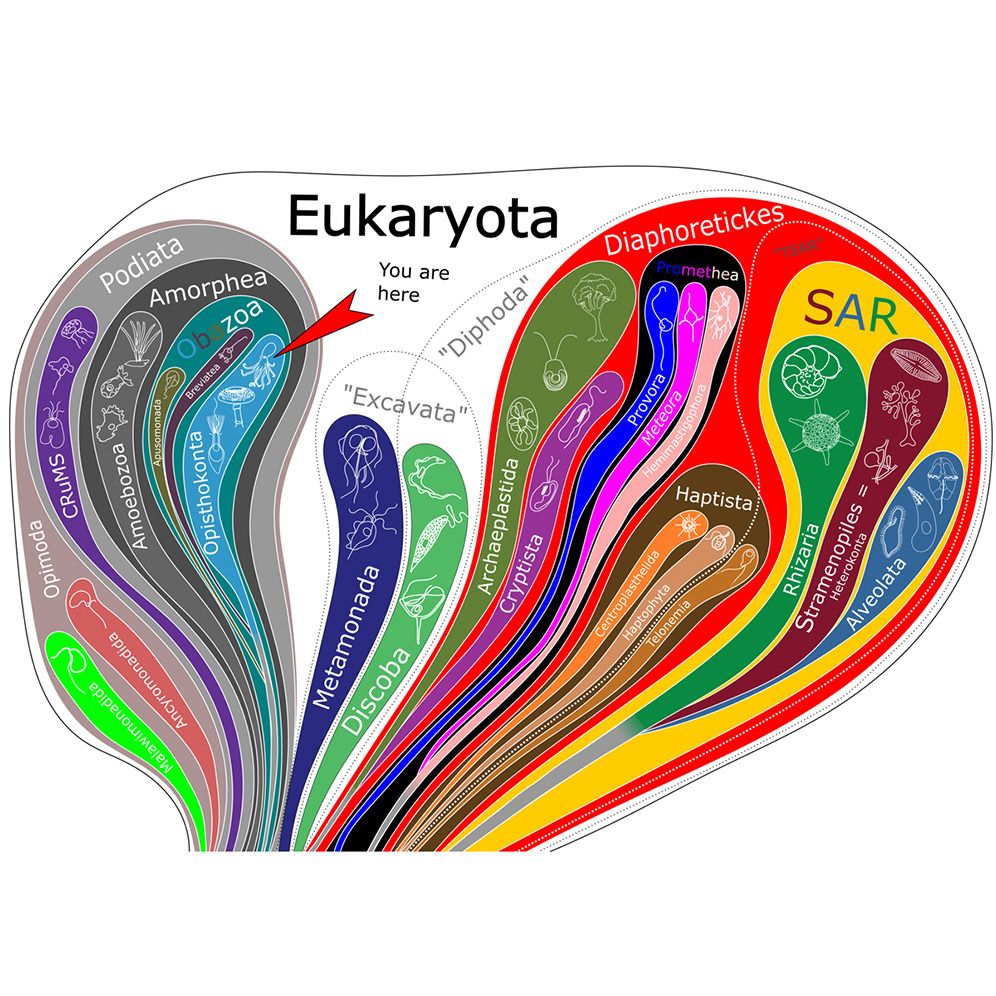
Promethea = PROvora + MEteora + HEmimastigophora. The new supergroup, unifying previously “orphan” lineages with gene-rich mitogenomes. Position of #telonemids is still uncertain. #protistsonsky tinyurl.com/yk9xkt49
24.07.2025 14:50
👍 44
🔁 22
💬 0
📌 1


eartharxiv.org/repository/v...
In our new preprint we describe in detail the software & workflows behind www.SewageMap.co.uk! One for the open environmental data nerds out there :) 💩 🧑💻 🏞️
31.07.2025 06:41
👍 16
🔁 7
💬 1
📌 0

Happy to have contributed to this great article: #Protist genomics: key to understanding eukaryotic evolution. Congrats Alexandra Schoenle et al. #ProtistsOnSky
authors.elsevier.com/sd/article/S...
13.06.2025 06:43
👍 87
🔁 42
💬 3
📌 5

Schematic representation of Eukfinder workflows. Eukfinder is a taxonomic classification-based bioinformatics approach to retrieve microbial eukaryotic nuclear and mitochondrial genomes from WGS metagenomic sequencing data. Eukfinder has two different workflows based on the input files. (a) Eukfinder_short utilizes Illumina short reads, and the first round of classification assigns reads into 1 of 5 distinct taxonomic categories (Archaeal, Bacterial, Viral, Eukaryotic, and Unknown). Next, Eukaryotic and Unknown reads are assembled into contigs which undergo a second round of classification to generate potential eukaryotic sequences. (b) Eukfinder_long uses assembled contigs or long-read sequencing data and only performs one round of classification to select Eukaryotic and Unknown contigs. The potential eukaryotic contigs can then be further separated into MAGs by a separate binning workflow to generate draft eukaryotic nuclear and mitochondrial genomes.
Introducing Eukfinder! A bioinformatics pipeline that identifies eukaryotic sequences from metagenomic data. The tool is valuable for reference-independent and cultivation-free studies of eukaryotic microbial genomes from environmental samples. #mBio: asm.social/2nl
23.04.2025 17:55
👍 16
🔁 9
💬 0
📌 1

🌊 New research reveals that coccolithophore blooms significantly enhance carbon export by increasing particle fluxes.
This finding underscores their vital role in oceanic carbon sequestration and has important implications for climate models. www.researchsquare.com/article/rs-6...
18.04.2025 22:36
👍 10
🔁 1
💬 0
📌 0
Eukfinder: a pipeline to retrieve microbial eukaryote genome sequences from metagenomic data journals.asm.org/doi/full/10.... #jcampubs
11.04.2025 21:23
👍 21
🔁 11
💬 0
📌 0

Hans Christian Andersen : The Drop of Water
Hans Christian Andersen: life and works - research, texts and information
We are glad to announce that #MidweekMicrobe is back!🥳 Today we are celebrating the International Children's Book Day coinciding with Hans Christian Andersen's birthday. A creative mind, Hans published the tale The Drop of Water in 1847, available at andersen.sdu.dk/vaerk/hersho....
02.04.2025 17:20
👍 7
🔁 4
💬 1
📌 0