

When I was in @hitenmadhani.bsky.social lab, we used a blend of lab-purified taq and Pfu to do colony PCRs on Cryotococcus that was robust. You can purchase Phusiom from the UCB MacroLab qb3.berkeley.edu/facility/qb3...

Published an op-ed for @cnn.com: “Nobel laureate: I owe America my success. Today, its scientific future is in danger.”
A personal reflection on what’s at stake as science funding gets slashed. I’d be grateful if you could amplify both in and beyond the science world.
www.cnn.com/2025/04/09/h...
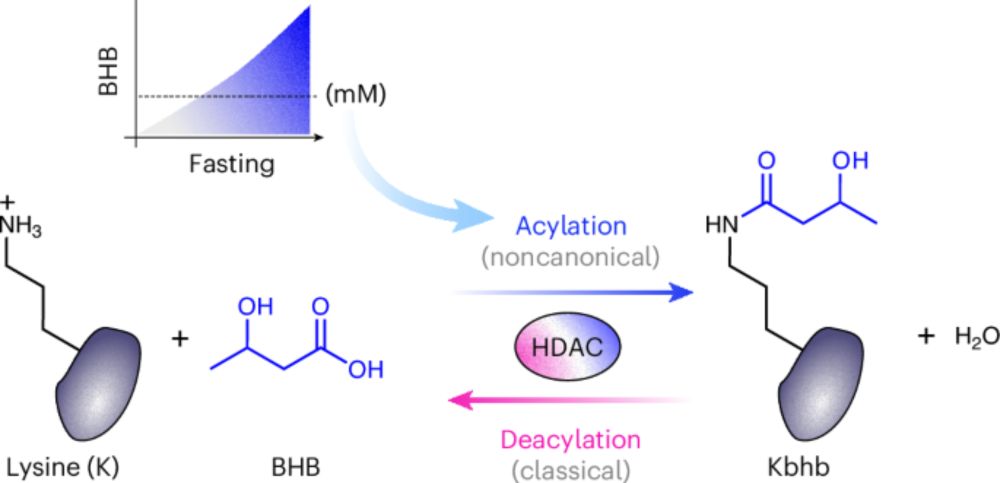
I could not be more excited to have our lab's first story online where we report our discovery that HDACs ~reverse~ their activity to ADD acyl groups to lysine! We found this for our favorite ketone body, BHB, but this pathway controls other lysine modifications too!🧵
www.nature.com/articles/s41...
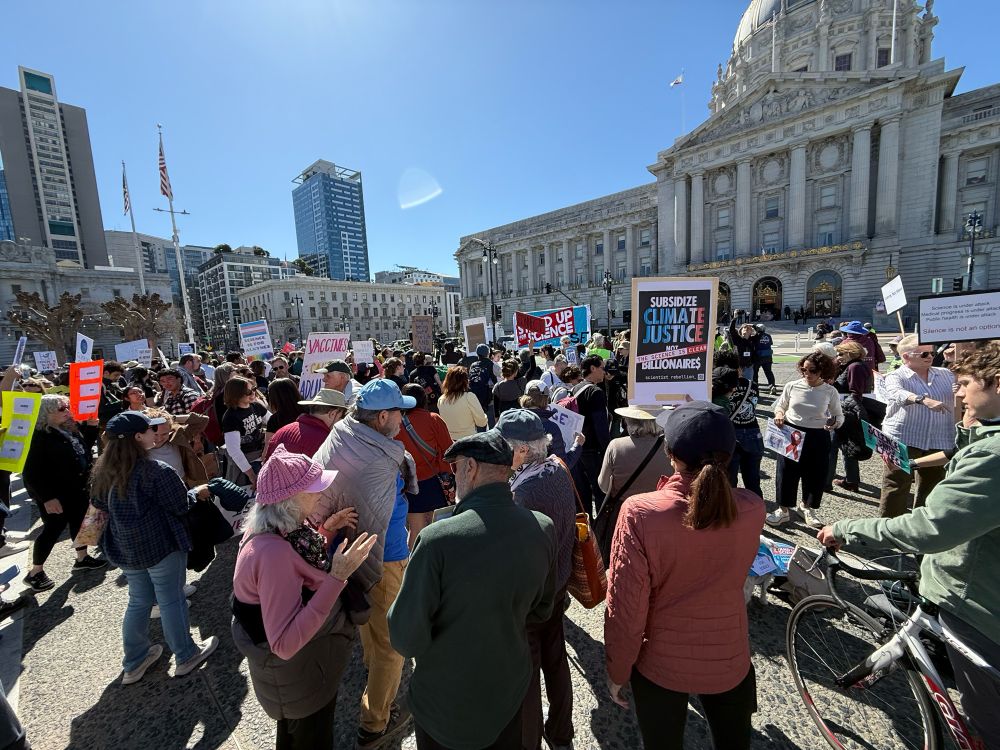
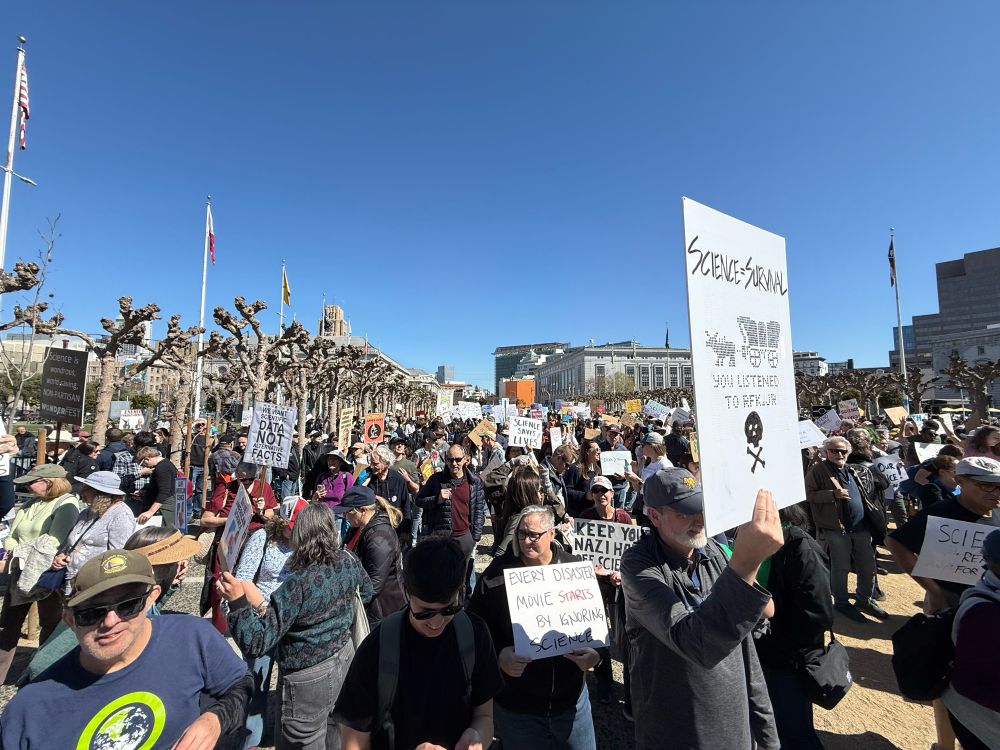


Awesome turnout @standupforscience.bsky.social SF today! The Oregon Trail inspired sign was one of my favorite!
Yup, but any of the current single cell antibody panels should work.
Hi John, we do think it's possible to capture protein information using tagged antibodies that have polyA barcodes or if using direct capture ADTs, spike in a splint oligo.

Welcome to the Bluesky account for Stand Up for Science 2025!
Keep an eye on this space for updates, event information, and ways to get involved. We can't wait to see everyone #standupforscience2025 on March 7th, both in DC and locations nationwide!
#scienceforall #sciencenotsilence
Just look at these paragraphs from the AP story.
Hegseth called Brown unqualified solely because he's Black. Then they fired him... and replaced him with a white guy so indisputably unqualified that he requires a Presidential waiver.
This is what "merit" means to them.
apnews.com/article/trum...
Can I give some unsolicited advice? When communicating on this catastrophe, don’t lead with “I know the NIH is inefficient and needs serious reform, but…”. You are undercutting the entire message and buying into the frame of the arsonists. This is not a scientist audience. No “to be fair” points
I’m reaching out to my vendors to push them to use their lobbying infrastructure to point out the devastating effects the NIH change will have on the USA’s R&D engine.
all y'all academics who have been naively convinced IDC only goes to useless sub deans are about to find out. they are going to claw the real costs of research out of your unchanged direct costs.
Indirect rates are not determined by NIH. Instead, they are the product of elaborate negotiations between each institution and the “cognizant federal agency” according to an Office of Management and Budget document, Circular A-21.
1/n
1. Today the NIH director issued a new directive slashing overhead rates to 15%.
I want to provide some context on what that means and why it matters.
grants.nih.gov/grants/guide...
Congratulations Chris!
No worries Andrew. Thank you for the responses, this info is helpful. Can’t wait to try DV on our future system!
Hi Andrew, we are spec’ing out a server. It seems DV utilizes extensions on Intel CPUs, so we’ll go with them. Any thoughts on the L4 vs L40S. Is there a possibility DV will require >16GB GPU memory in the near future?
Thanks!
We only upgraded one of our systems as a “just in case”. Our update required an engineer visit so that might also play a role?
We are currently offering 10X GEX lane queues (28x10x10x90). If there is enough interest for a PE50/10X ATAC (51x12x24x51), we will start that as well.
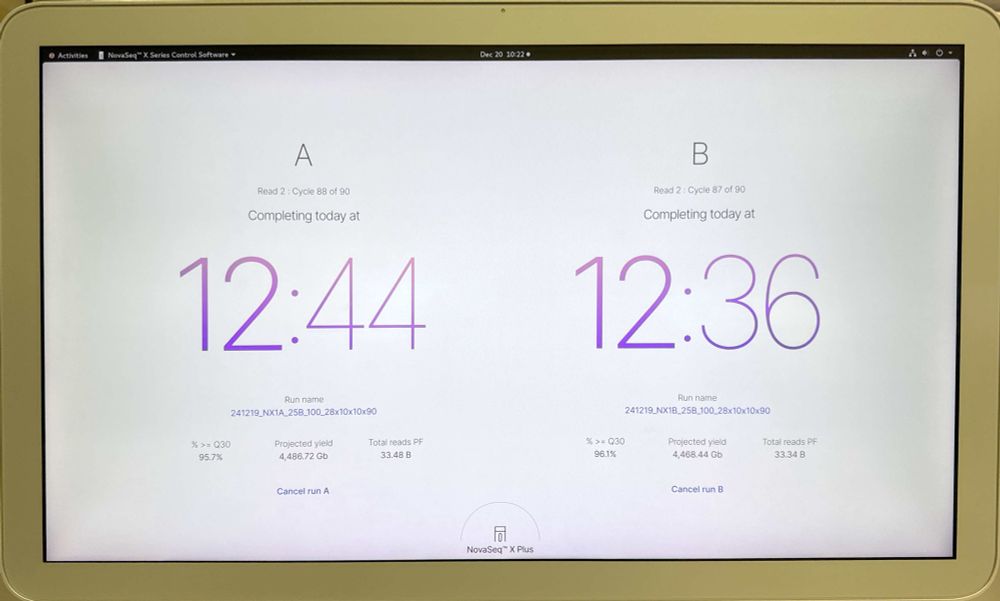
We just upgraded one of our Illumina NovaSeqX’s to enable the new 25B 100 and 200 cycle kits. New software is supposed to slightly raise PF% and we are seeing that. 10X GEX libraries previously generated 32B reads per flow cell. We seeing over 33B, almost 67B reads in 24 hours for a dual FC run!
Some preps recommend different cycles (10X SC and for Visium a qPCR titration). With clinical metagenomic samples, we are not comparing across samples so no worries there.
We’ll look at for bulk-RNA-seq experiments but even with constant cycles, the n6 will give you quant and pooling info.
UCSF users, we will have a training to be scheduled the second week of January. Details will go out on our listserve and posted on our website.
The iconPCR solves these issues. Samples will be within 2-fold concentration of each other and the software uses the fluorescence level to calculate relative concentrations and pooling volumes to further normalize pools. You get a pool that’s ready to go on your production scale sequencing runs.
This works fairly well but adds time and cost and some samples are amplified too little, and others too much, resulting in adapter dimer carryover in bubble products.
We see many applications but our first one will be metagenomics from clinical samples where inputs vary widely. To Amie this high throughput, we have been pooling and equal volume of many samples after PCR, doing a cleanup on the pool, running a MiSeq nano, and repooling based on read distribution.
The system will place each sample at a low temp hold once they have reached a user-defined set point. If anyone used the old Kapa Real Time NGS amp kit, this allows you to do this automatically with your choice of PCR mix. Just add a little bit of SYBR Green.
We just got our n6 Tec iconPCR installed. The system monitors NGS library amp with real time functions but each of the 96 wells is independently temperature-controlled and monitored. This is super helpful when you have varying inputs. www.n6tec.com
The new MiSeq i100 should be good to check pooling for the NovaSeqX. Catherine's group is seeing good consistency between the two platforms, but from first glance, it does appear the MiSeq i100 is more consistent with the older iSeq (blue and orange lines).
Hi Andrew, any benchmarking on different GPUs? Particularly interested comparing the L40S and L4. Thanks!
150pM by our custom ddPCR assay we’ve been using for the past decade. Taking loading concentration with a grain of salt - as you know, different methods and users can result in different quants, especially our assay which uses a 10^7 dilution.