The scientific part of the event starts with a talk by @sergiocruzleon.bsky.social on bridging in-situ cry-ET and multi scale simulations
The scientific part of the event starts with a talk by @sergiocruzleon.bsky.social on bridging in-situ cry-ET and multi scale simulations
We resolved the molecular architecture of a condensate—inside cells. 🤯 By combining in situ cryoET with multiscale simulations, we show how a single point mutation modifies molecular interactions, shifts the condensate’s material properties, and alters its biological function. Preprint 👇
It’s out! Using cryo-ET in Dicty cells, we take a fresh in situ look at vaults. Surprisingly, we uncover vaults associated with ER and NE membranes, and find that many vaults enclose ribosomes in defined orientations, opening new avenues to their cellular function! www.biorxiv.org/lookup/conte...
We developed a 'smart STED microscope' to detect rare objects/events in cells, and unraveled the 3D morphology of a nuclear membraneless organelle (paraspeckle). Amazing collaboration with Sjoerd Stallinga, Bernd Rieger (TU Delft) and @mixmue.bsky.social 🙏 (1/3) www.nature.com/articles/s41...
Just published! We share an open-source workflow to measure membrane thickness from tomograms, including a tutorial with 3D visualizations. We analyze thickness variations across organelles and reflect on where to define a membrane boundary. @becklab.bsky.social #teamtomo rupress.org/jcb/article/...

Structure of the ribosome-bound SND3 translocon complex from the heat-tolerant fungus Chaetomium thermophilum. The SND3 translocon inserts proteins into the endoplasmic reticulum membrane as they are produced by the ribosome.
SNDing proteins into the membrane! Our new publication from @melaniemcdowell.bsky.social ’s group identifies the SND3 protein as a new route for membrane protein insertion! 🍄 📘 Read more here: www.mpg.de/25599408/102... Image: Louise Duever.

Deep learning-based interpolation using cryoTIGER developed by the authors in this new Communications Biology manuscript enhances cryoET reconstructions. @mpibp.bsky.social
@becklab.bsky.social
@jpkreysing.bsky.social
@sergiocruzleon.bsky.social
doi.org/10.1038/s420...

Photo of Sergio Cruz and text: Sergio Cruz-León receives the 2026 Theory & Computation Postdoctoral Research Award
🎉 Congratulations to our postdoc @sergiocruzleon.bsky.social for receiving the 2026 Theory & Computation Postdoctoral Research Award from the Biophysical Society! 🎉 Sergio will present his work at the Theory & Computation subgroup Symposium during the 2026 Biophysical Society meeting.

Ready to lead pioneering research that bridges systems-level investigations of biological systems to molecular mechanism? The EMBL Molecular Systems Biology Unit in Heidelberg is hiring a Group Leader!
That's a wrap! The results of the first #cryoEM heterogeneity challenge are up on biorxiv!
biorxiv.org/content/10.110
Proud to share our first lab pre-print: “SND3 is the membrane insertase within a fungal multipass translocon” where @tzujingyang.bsky.social solved the structure of a ribosome-associated SND3-translocon complex involved in ER membrane protein insertion ➡️ doi.org/10.1101/2025...
Thanks 😀🧬
🧬😄! Thanks!
😄 thank you!
😄 thanks!
Thanks!
Thanks!
Thanks!
Thanks!
New preprint on 3D heterochromatin architecture in human cells! Great collab with @sergiocruzleon.bsky.social & @johannesbetz.bsky.social from @hummerlab.bsky.social, @marinalusic.bsky.social & the Turoňová lab. Many thanks to my supervisor @becklab.bsky.social. bioRxiv: tinyurl.com/3a74uanv 🧵👇
Excited to share our preprint on the molecular architecture of heterochromatin in human cells 🧬🔬w/ @jpkreysing.bsky.social, @johannesbetz.bsky.social,
@marinalusic.bsky.social, Turoňová lab, @hummerlab.bsky.social @becklab.bsky.social @mpibp.bsky.social
🔗 Preprint here tinyurl.com/3a74uanv

Our paper on prediction of phase-separation propensities of disordered proteins from sequence is now published:
www.pnas.org/doi/10.1073/...
The paper has been substantially updated compared to the preprint including new experimental data and using the neural network to finetune CALVADOS. 1/n
✨✨ It's #glycotime for #HIV Env ✨✨
N-linked glycans modulate flexibility & MPER epitope exposure #glycotime
Huge effort by @shehata92.bsky.social @lcasalino88.bsky.social with cryoET of Env in VLPs by @thevillalab.bsky.social & team 💪
Would love feedback!
www.biorxiv.org/content/10.1...

Job alert: Join us in Mainz as
Max Planck Research Group Leader (W2) in Molecular Design
...and make your own research dreams happen on de novo design, generative models, proteins, materials ...
tinyurl.com/r2xjxnuk
@mpip-mainz.mpg.de
Thrilled to share these experiments on shaping focused ion beams to explore different ways of milling. Here, we explore using different beam geometries to generate cellular thin sections/lamellae. The video shows elongating the beam on a charging spot burn www.biorxiv.org/content/10.1...
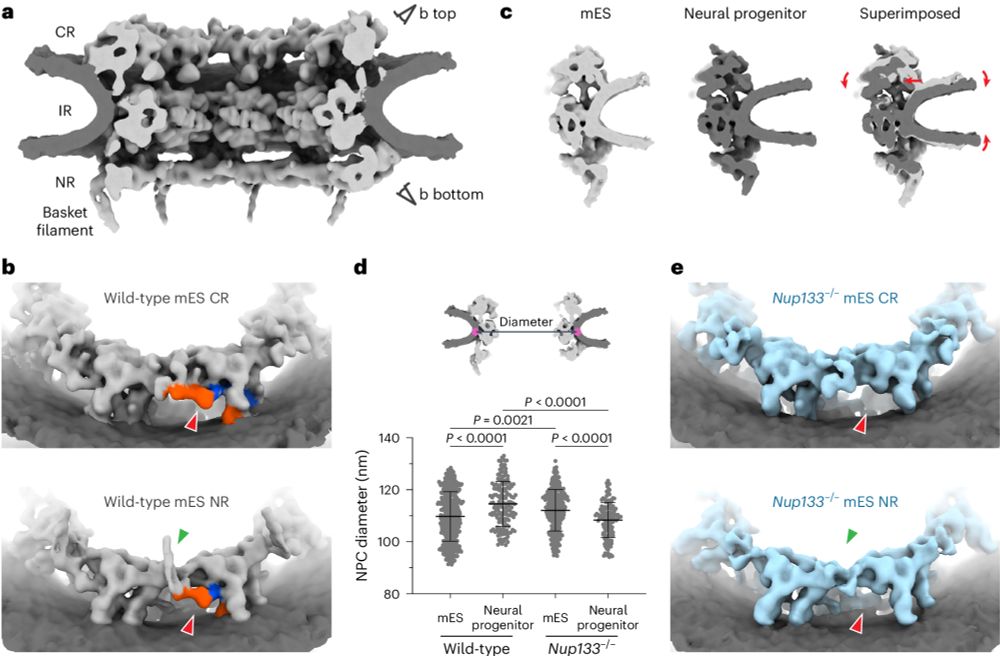
Excited to share our latest work on nuclear pore complex scaffold integrity and its perturbation. Now out in Nature Cell Biology: rdcu.be/eg4i9
Great work by Reiya Taniguchi who just opened his own research group back in Japan! 🥳
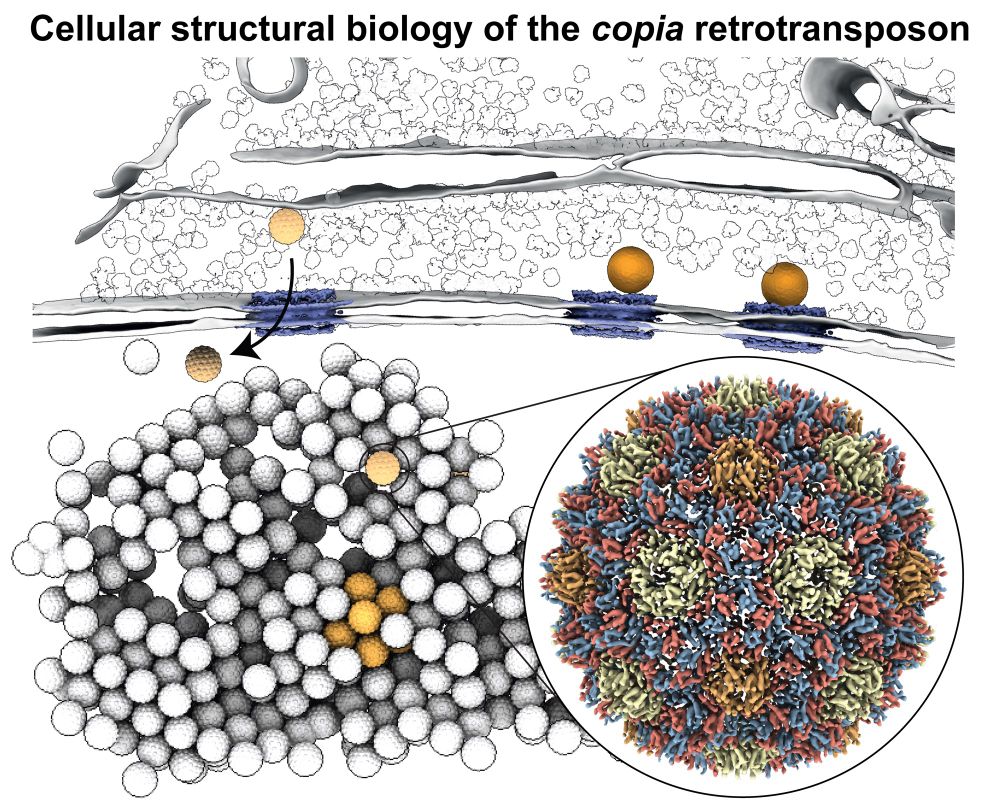
Happy to share our manuscript on the in situ visualization of the copia retrotransposon in its final form today published in @cellcellpress.bsky.social www.cell.com/cell/fulltex.... What’s new?
Interested in #RNA modelling across multiple scales? Join us for the @cecamevents.bsky.social meeting at SISSA, Trieste, May 19-22! www.cecam.org/workshop-det... We still have slots for contributed talks. Deadline March 23. Co-organized with @magistratolab.bsky.social and @marcodevivo.bsky.social

🤩 Highly optimized software running on a SOA #HPC cluster provides sufficient #computing power to #simulate #biomolecular systems at the mesoscale of viruses and organelles, and potentially small cells in the near future... Want to know more? Check out our paper 📑 in #JCC 🔗 doi.org/10.1002/jcc....

Excited to see work led by @autophazeke.bsky.social, @minghaochen.bsky.social, Thanh Nguyen, and Ainara Claveras out in @science.org today, the culmination of a 15-year project to understand structurally how the synthesis of the critical lipid of autophagy, PI3P, is regulated. tinyurl.com/ywvjh8u4