Courtesy of @martibartfast.bsky.social , we have a new release of AllTheBacteria which adds another 322,920 assemblies, covering all ENA (illumina, isolate) prokaryotes to May 2025.
allthebacteria.readthedocs.io/en/latest/ov...
Courtesy of @martibartfast.bsky.social , we have a new release of AllTheBacteria which adds another 322,920 assemblies, covering all ENA (illumina, isolate) prokaryotes to May 2025.
allthebacteria.readthedocs.io/en/latest/ov...
It's a bit last minute, but we have an RA/junior postdoc position available in my lab (closing Sunday). Please reach out if you're interested in developing and applying methods to analyse microbial genomes and metagenomes! careers.petermac.org/job/MELBOURN...

To make sure the whole Shigella community will be gathered at this inaugural meeting, we have re-opened abstract submission for poster presentation until the end of the month. Please RT!
www.shigella2026.conferences-pasteur.org/registration...

Spotted on the Chappell Roan merch store. I don’t think they know what Shigella is… what am I missing???

Our new paper on Insertion Sequences (IS) in #Klebsiella
- Lineages have vastly different IS loads and profiles
- An inverse relationship between IS load and metabolic capacity, in particular phosphorus use, consistent with early reductive evolution.
www.microbiologyresearch.org/content/jour...

Finally got my #triplej #hottest100 votes in! Thought I heard no new music this year but that was a lie, my short list was huge. Ur A Rat and Keith I can never get out of my head
(1/7) Really pleased to have this published in @natcomms.nature.com
We looked at all public genomes encoding blaIMP (carbapenem #AMR ) 🦠🧫🧬💊. We show 5 IMP variants achieved global endemicity while 2 are regionally endemic.
www.nature.com/articles/s41...
#MicroSky #microbiology #IDSky #EpiSky 🧪

Absolutely stoked to have this published in @plosbiology.org
We looked at the metabolism of #Klebsiella pneumoniae 🦠🧫. We not only demonstrated lineage-specific #metabolism, but that lineages can cross-feed and support each other.
journals.plos.org/plosbiology/...
#MicroSky #microbiology 🧬 🧪 💊
The GTDB website now has an ANI calculator based on skani that supports uploading of user genomes. Try it at gtdb.ecogenomic.org/tools/skani.
Find more information about @jimshaw.bsky.social fantastic tool at www.nature.com/articles/s41....
A new tool we’ve developed to identify AMR-associated SNPs in short- and long-read metagenome data….allows a greater understanding of the total resistome of a sample
I’m so jelly I missed it! Any plans to make some of the materials available?
I’ve been enjoying seeing the updates, wishing I’d made the trek to Adelaide now!
I can’t explain why but reading the news feels like this at the moment
Very interesting to see what amendments were pushed by different parties for this bit of legislation. The One Nation amendment to remove a focus on public health issues caused by climate change (thankfully voted down) was typically 🙄
New online! Microbial genomics for antimicrobial resistance ecology and action

Are you hungry for some Shigella news? Why not come to the 1st International Shigella meeting www.shigella2026.conferences-pasteur.org/home 20-24 April 2026 Paris

There are millions of openly available microbial genomes, but searching them can be slow.
Until now 🥁
Introducing LexicMap, a new alignment tool that lets scientists search these data in minutes, helping track antibiotic resistance, trace outbreaks, and more.
www.ebi.ac.uk/about/news/r...
🦠

Happy to share that the paper describing Autocycler is now 100% up:
doi.org/10.1093/bioi...
(1/3)
To summarise our recent pre-print: Autocycler, the automated consensus assembler, when used with Nanopore long-read only Enterobacterales assemblies, produces more complete chromosomes and plasmids, with an accuracy comparable to hybrid assemblies.
A great opportunity to attend the #Shigella meeting April 20-24, 2026! Abstract submissions are open! @shigellameeting.bsky.social #Microsky

Come and join us at @shigellameeting.bsky.social in Paris 2026. Registrations are now open! www.shigella2026.conferences-pasteur.org/home and follow the Shigella starter pack go.bsky.app/6Vbwhjc (message to be added)! 215 days to go!

I am beyond excited to announce that ggplot2 4.0.0 has just landed on CRAN.
It's not every day we have a new major #ggplot2 release but it is a fitting 18 year birthday present for the package.
Get an overview of the release in this blog post and be on the lookout for more in-depth posts #rstats
Academia may not give you job security, flexibility, or wealth, but it will let you unexpectedly connect to eduroam in foreign cities
Really liked the explanation about human understanding, and what we miss when we use black box models. This has always been my gripe with prediction models for eg AMR - I want to know why that feature is associated with resistance!

Register today for the 14th International Meeting on Microbial Epidemiological Markers (IMMEM XIV)! Topics range from how genomics inform resistance profiling to pathogen surveillance in a #OneHealth framework. 🧬
✍️ Register: ow.ly/gjOZ50WpOXX
👀 Programme: ow.ly/Whj850WpOXY
#IDSky
This surprisingly relaxing footage is from SIX MILES under the ocean – and it’s the deepest ecosystem yet discovered
Huge amount of work in this paper, very proud of Hugh getting through all these benchmarking tests. We’re much more confident now using ONT-only data for our clinical questions
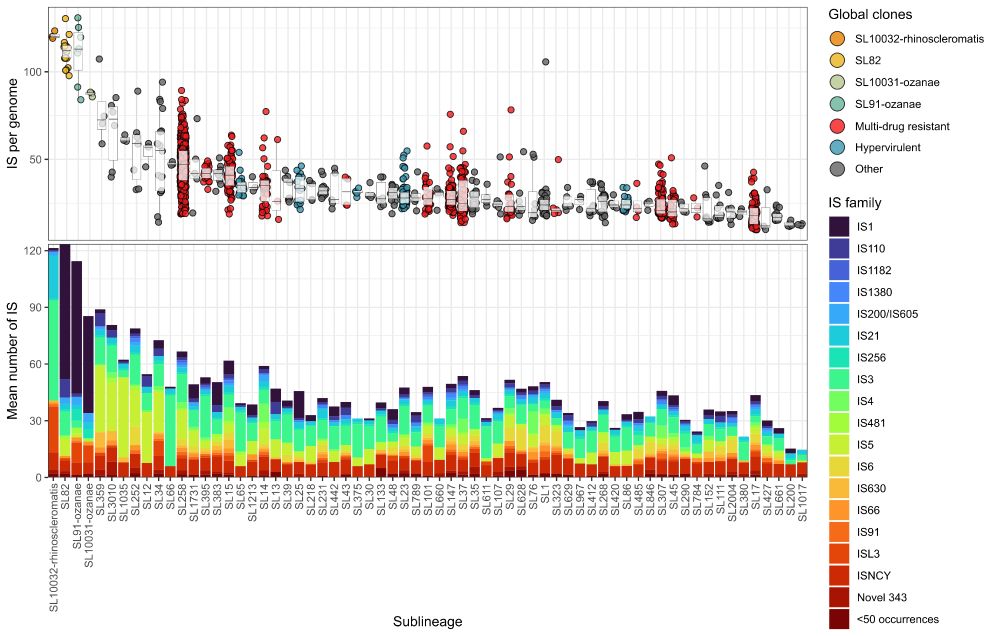
Graph showing IS loads by Klebsiella sub-lineage
New preprint where we analysed IS loads in 🦠 #Klebsiella pneumoniae lineages and found they're inversely associated with nutrient usage. We propose an insertional tolerance model, whereby IS can only insert into metabolically-tolerable sites.
#IDSky #MicroSky 🧪
www.biorxiv.org/content/10.1...

Fold first, ask later: structure-informed function annotation of Pseudomonas phage proteins
Structural predictions to improve #phage genome annotation!
@hannelorelongin.bsky.social @gbouras13.bsky.social y.social @susiegriggo.bsky.social @veravannoort.bsky.social
www.biorxiv.org/content/10.1...

Registration and abstract submission for MicroSeq2025 is open. Abstract submission closes August 8, 2025. Participants can register at microseqconference.com. MicroSeq2025 is an online meeting being held on September 3rd and 4th, focused on research being conducted on sequencing data generated from microbiological samples (bacteria, fungi, and viruses). MicroSeq2025 is supported by The Australian Society for Microbiology, the Australasian Virology Society, and Microbial Genomics.
🚨Abstract submission & registration for #MicroSeq2025 is OPEN! 🚨Get in quick! The first 50 registrations are FREE for PhD students and ECRs who are current ASM members, thanks to @aussocmic.bsky.social.
Head to our website to register now 👇
www.microseqconference.com/registration...