Very happy that MuViT, our (CVPR 2026!) work on learning across spatial scales in microscopy w transformers, is out! Check Martin's thread for a nice walkthrough :)
Big thanks to my advisors @maweigert.bsky.social @gioelelamanno.bsky.social!
🖥️: github.com/weigertlab/muvit
📜: arxiv.org/abs/2602.24222
02.03.2026 14:59
👍 19
🔁 7
💬 5
📌 0

It was super cool to hear @alex-krull.bsky.social talking about his research at @humantechnopole.bsky.social. Alex never fails to inspire his audience 🫠🤩
11.02.2026 10:46
👍 10
🔁 1
💬 0
📌 0
Great work by our colleagues
@afoix.bsky.social
@virginieuhlmann.bsky.social
@alex-krull.bsky.social
As presented at this year’s @neuripsconf.bsky.social
18.12.2025 08:57
👍 2
🔁 1
💬 0
📌 0
If you are at @neuripsconf.bsky.social and interested in shapes, check out Anna's work! Thank you @afoix.bsky.social! It was a real pleasure working on this with you and @virginieuhlmann.bsky.social and I am sad I can't join in San Diego.
01.12.2025 20:56
👍 4
🔁 0
💬 0
📌 0
We’ll be at @neuripsconf.bsky.social on Wednesday, 3 December, from 11 a.m. to 2 p.m. (San Diego time). Come and chat with us about why distance matrices and invariances are cool! 🎉 #InterdisciplinaryML #neurips2025 #bioimageanalysis #shapequantification
01.12.2025 08:27
👍 23
🔁 3
💬 0
📌 2
Cheers, @afoix.bsky.social. Thank you all of you for submitting, reviewing, organising and attending! Thank you #ICCV2025 for hosting us. Also thank you @unibirmingham.bsky.social for sponsoring #BIC again this year!
17.10.2025 11:50
👍 3
🔁 0
💬 1
📌 0
Thank you, @afoix.bsky.social!
I am really happy I could be part of this one. Everybody, please enjoy the new ShapeEmbeding goodness!
17.10.2025 11:43
👍 4
🔁 1
💬 0
📌 0
Thank you @afoix.bsky.social , thank you @virginieuhlmann.bsky.social! I am very happy and proud I was part of this one!
23.09.2025 09:05
👍 7
🔁 0
💬 1
📌 0
Cheers, for sharing, @paveltomancak.bsky.social! What a brilliant example for rotation-invariance or the lack of it.
Will go into the lecture.
21.08.2025 14:58
👍 1
🔁 0
💬 0
📌 0
Sept 18: Learn how to remove noise from imaging data with supervised, self-supervised & generative AI. With @alex-krull.bsky.social (University Birmingham).
Register for the series 👉 bit.ly/6-image-proc...
@helmholtz.de
#imaging #ImageDenoising #AI
18.08.2025 07:30
👍 1
🔁 2
💬 0
📌 0
Looking forward to my talk at heidelberg.ai. I will touch on topics like #MapFreeReloc and #ACEZero, i.e. how to derive scene representations from very few (one!) or very many images.
04.07.2025 07:08
👍 14
🔁 1
💬 1
📌 0

17.07.2025 17:48
👍 1
🔁 0
💬 0
📌 0

Please meet Dr. @stonks1.bsky.social!
Couldn’t be more proud!
15.07.2025 17:04
👍 4
🔁 1
💬 0
📌 0
Submit your method to our denoising challenge!!! 👍
14.07.2025 09:19
👍 7
🔁 3
💬 0
📌 0
Vacancy ID 12244
Do you like developing new AI vision methods for microscopy image analysis? You love theory & implementation? 1 week left to apply for a fully funded PhD position in our lab in Dresden 🇩🇪! Topics: object detection/tracking, multimodal models & more. DM/email for details! #PhD #AcademicJobs #GPUsgoBrr
10.07.2025 13:49
👍 25
🔁 20
💬 1
📌 4
Rock on, Pavel! 👍👍👍
09.07.2025 19:33
👍 1
🔁 0
💬 1
📌 0

The 2nd AI4Life Challenge is live!
Calling the AI & bioimaging community to tackle a key microscopy challenge: removing noise while preserving detail.
📦 Paired noisy/clean datasets
📈 Ground-truth evaluation
🧠 DL focus
Build, test, compete 👉 ai4life.eurobioimaging.eu/challenge-2/
27.06.2025 13:33
👍 27
🔁 15
💬 1
📌 2
Join the challenge, get the data, contribute your solution!!! 🎉
30.06.2025 08:55
👍 5
🔁 2
💬 0
📌 0

BioImage Computing
a truly interdisciplinary workshop
The submission for 2025's BioImage Computing @iccv.bsky.social is now open. If things go like the last years, I am looking forward to all you brilliant submissions.
www.bioimagecomputing.com
10.06.2025 21:40
👍 5
🔁 2
💬 0
📌 0
Congrats, guys!
06.06.2025 22:36
👍 4
🔁 0
💬 0
📌 0
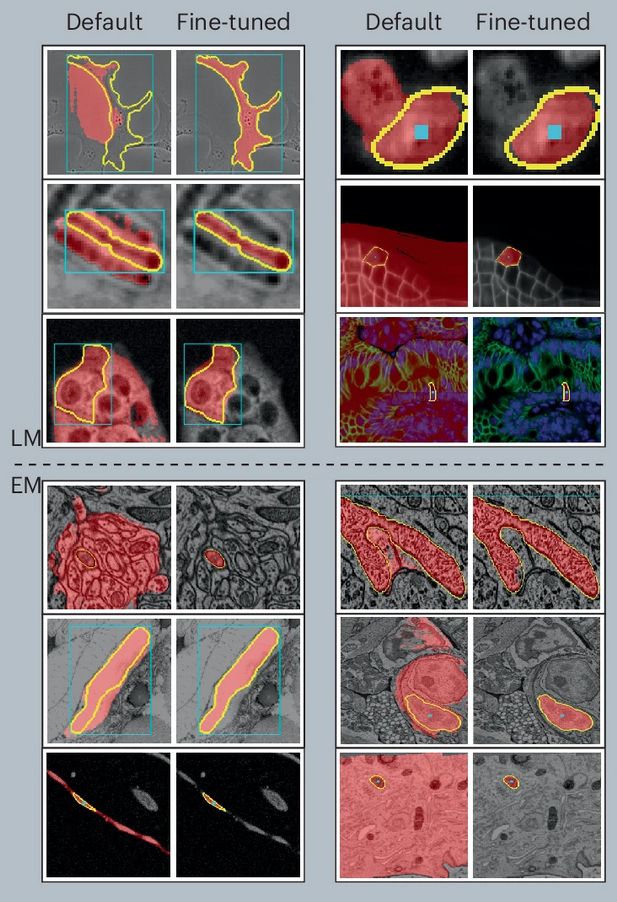
Improvements in LM (top) and EM (bottom) of our micro-sam model (finetuned) compared to the default SAM model.
After a long journey, Segment Anything for Microscopy is now published in Nature Methods! We significantly improve SAM for interactive and automatic segmentation in light and electron microscopy and build a user-friendly tool.
www.nature.com/articles/s41...
12.02.2025 11:41
👍 352
🔁 121
💬 11
📌 7

Tweedie's formula is super important in diffusion models & is also one of the cornerstones of empirical Bayes methods.
Given how easy it is to derive, it's surprising how recently it was discovered ('50s). It was published a while later when Tweedie wrote Stein about it
1/n
18.03.2025 06:12
👍 65
🔁 14
💬 1
📌 0
Original post on social.tchncs.de
"Having trouble selling your #Tesla? Tired of being associated with a toxic, Nazi-adjacent brand? We understand that there’s a shrinking market of buyers and limited selling options for current owners.
Our experienced team specializes in discreetly acquiring and transporting your car to […]
13.03.2025 14:39
👍 0
🔁 3
💬 1
📌 0
Wow! Congrats!
13.03.2025 11:14
👍 2
🔁 0
💬 0
📌 0
@docmilanfar.bsky.social , love this. Max likelihood points to x. This refers to the fact that the noisy data x is actually the max likelihood est. of u?
04.03.2025 08:29
👍 0
🔁 0
💬 1
📌 0

Have you said thank you once to Tweedie?
04.03.2025 05:58
👍 37
🔁 4
💬 2
📌 0
🚨 Fantastic opportunity to conduct a PhD in Europe! European funding and fantastic international community! 👩🎓
24.02.2025 13:31
👍 7
🔁 5
💬 0
📌 0

Poster for Theory of Living Matter seminar series talk 05 March 2025 at 16:00 UK time. Speaker is Virginie Uhlmann from BioVisionCenter, University of Zurich and EMBL-EBI. The talk is entitled "Turning morphology into numbers across microscopy scales and modalities", with the following abstract: Microscopy image data constitute the primary source of observational data on living systems across scales, and recent advances in microscopy technologies now enable the multi-modal observation of identical biological specimen. A key question emerges from this abundance of image data: how can we consistently quantify what we see? This endeavour is particularly complex for morphology, which involves intricate combinations of visual features that appear fundamentally different across various imaging modalities. Motivated by this challenge, my research group has been focusing on abstracting the problem of image-based morphology quantification from specific imaging modalities. In this seminar, I will present how we are building bits by bits the computational tools needed to extract modality-agnostic shape descriptors from microscopy images, how we use them to investigate fundamental biological questions, and what are the promising directions we are pursuing towards establishing a general framework to quantify biological morphology across imaging modalities.
🚨Next week, Wed 05/03 | 16:00 UK - Online Seminar🚨
"Turning morphology into numbers across microscopy scales and modalities" by Dr Virginie Uhlmann.
To attend online please register to our 📧 for the zoom link: lists.cam.ac.uk/sympa/subscr...
27.02.2025 16:06
👍 17
🔁 10
💬 0
📌 2

Pflanzenzellen, die mit einem Fluoreszenzmikroskop aufgenommen und mit dem Modell automatisch segmentiert wurden. Die zugrunde liegenden Daten sind dreidimensional und das Bild zeigt eine Darstellung der segmentierten Zellen, die jeweils durch eine andere Farbe repräsentiert werden.
Foto: Nature Methods: 10.1038/s41592-024-02580-4

Segmentierung von Zellen in der Lichtmikroskopie mit μSAM. Das Bild zeigt, wie Zellen in der Phasenkontrastmikroskopie mit μSAM segmentiert werden können. Grüne Punkte und Kästchen zeigen die Benutzereingabe und farbige Masken die entsprechende Vorhersage des Modells.
Foto: Erstellt von Anwai Archit mit dem μSAM-Tool, verfügbar in Nature Methods: 10.1038/s41592-024-02580-4
Automatische Zellanalyse mit #KI: Forschende trainierten eine bestehende, KI-basierte Software neu. Das Modell „Segment Anything for Microscopy“ kann Bilder von Geweben, Zellen und anderen Strukturen genau segmentieren: s.gwdg.de/HqkMz2; s.gwdg.de/iTejKW
Forschungsteam mit bsky.app/profile/cppa...
26.02.2025 10:41
👍 11
🔁 3
💬 0
📌 0
Loved it!
25.02.2025 14:29
👍 1
🔁 0
💬 0
📌 0











