
The final piece of my PhD work is now published in Science Translational Medicine! We present a new framework to jointly infer epidemiological and antigenic parameters from multi-pathogen population serological studies🦠
www.science.org/stoken/autho...
18.12.2025 10:04
👍 32
🔁 13
💬 1
📌 1
For the first time, we found phylogenetic evidence of importations and local transmission of Macrolide-resistant #pertussis strains in France. Needs further monitoring!
Great work led by Valérie Bouchez, Carla Rodrigues and @sylvainbrisse.bsky.social!
Paper: shorturl.at/2Npmm
🦠 🧪 🛟
03.07.2025 08:30
👍 2
🔁 0
💬 0
📌 0

Fine-scale patterns of SARS-CoV-2 spread from identical pathogen sequences - Nature
The analysis of pairs of identical SARS-CoV-2 genome sequences enables characterization of transmission patterns between geographies and age groups.
Pathogen genomes can provide insights into underlying disease transmission patterns but new methods are needed to analyze large genome datasets. Our work using identical pathogen sequences to characterize fine scale SARS-CoV-2 transmission was just published in @nature.com tinyurl.com/bdzk9xjj 🥳
05.03.2025 16:48
👍 41
🔁 17
💬 2
📌 2
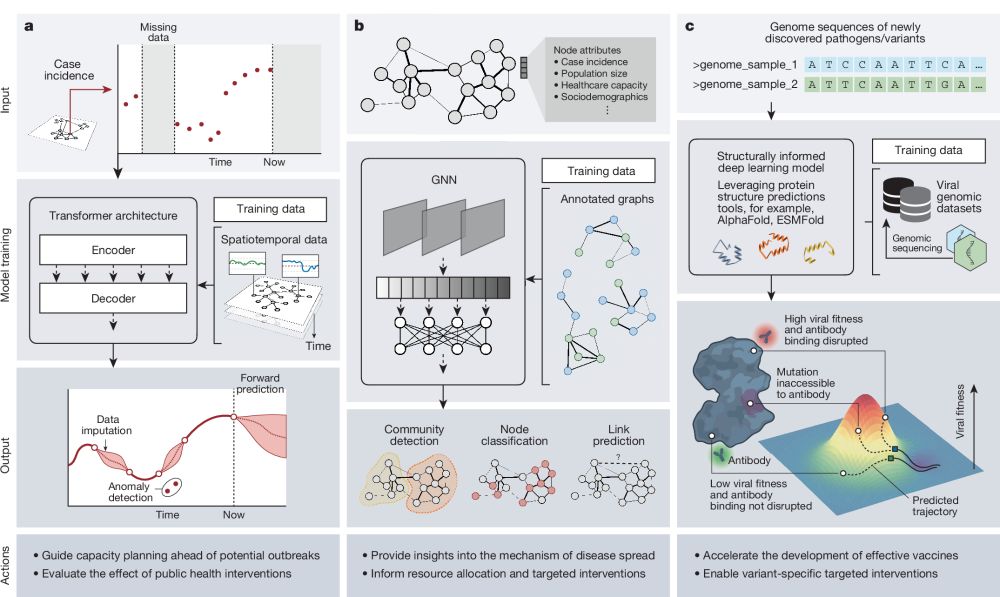
Artificial intelligence for modelling infectious disease epidemics
Nature - This Perspective considers the application to infectious disease modelling of AI systems that combine machine learning, computational statistics, information retrieval and data science.
AI is poised to accelerate understanding in infectious diseases, but its value needs to be demonstrated through close collaboration between research, industry, society, and policy.
Paper free to read: rdcu.be/eaxEw
Summary here: www.ox.ac.uk/news/2025-02...
20.02.2025 10:13
👍 69
🔁 31
💬 3
📌 5
Thanks!
31.01.2025 06:54
👍 0
🔁 0
💬 0
📌 0
Yes, but the method will even go beyond simply MDR/non-MDR and find what precise clades are spreading differently in your population. That's exactly what we did in the paper for the dataset of Samara, Russia.
09.01.2025 08:32
👍 1
🔁 0
💬 0
📌 0
Learning the fitness dynamics of pathogens from phylogenies - Nature
Phylowave, an innovative phylogenetic approach, can identify the main circulating pathogen lineages with increased fitness and the associated genetic changes, enabling the timely identification of…
Incredibly excited to share that our manuscript was just published in @nature.com ! What a way to start the new year! 🎉
https://buff.ly/4gyYCzx
We present phylowave, a framework that enables to learn the fitness dynamics of pathogens from phylogenies.
🧵 A thread... 1/n
#IDSky #IDModelling
02.01.2025 08:03
👍 215
🔁 87
💬 3
📌 4

Learning the fitness dynamics of pathogens from phylogenies - Nature
Phylowave, an innovative phylogenetic approach, can identify the main circulating pathogen lineages with increased fitness and the associated genetic changes, enabling the timely identification of eme...
🦠🔍 Tracking Pathogen Evolution in Real Time
A new tool, 'phylowave', has been developed to help scientists track how viruses and bacteria evolve. It identifies new variants with increased fitness, offering insights into diseases like #COVID19 and flu.
🔗 www.nature.com/articles/s41...
#SciComm 🧪
03.01.2025 05:59
👍 129
🔁 43
💬 1
📌 1
I am now working @ethzurich.bsky.social @ETH_BSSE with @tanjastadler.bsky.social and @neher.io with an ETH fellowship to develop more phylodynamic frameworks for large-scale datasets. Very excited to start this new year! 13/n
02.01.2025 08:04
👍 5
🔁 0
💬 1
📌 0
Developing this was a journey! Many thanks to the people who helped me along the way! Loréna Duret, super master student, my supervisors @hsalje.bsky.social and @julianparkhill.bsky.social and colleagues at Institut Pasteur @sylvainbrisse.bsky.social Valérie Bouchez. 12/n
02.01.2025 08:04
👍 6
🔁 1
💬 1
📌 0

GitHub - noemielefrancq/Phylowave_Learning-fitness-dynamics-pathogens-in-phylogenies: Code for the paper "Learning the fitness dynamics of pathogens from phylogenies"
Code for the paper "Learning the fitness dynamics of pathogens from phylogenies" - noemielefrancq/Phylowave_Learning-fitness-dynamics-pathogens-in-phylogenies
Phylowave code is available on GitHub. While it is not packaged, it comes with a full readme and description of best practices. All you need to start is R and a timed phylogeny. If you have any questions or suggestions, feel free to open an issue/drop me a message! 11/n
https://buff.ly/3PEj32l
02.01.2025 08:04
👍 3
🔁 0
💬 1
📌 0
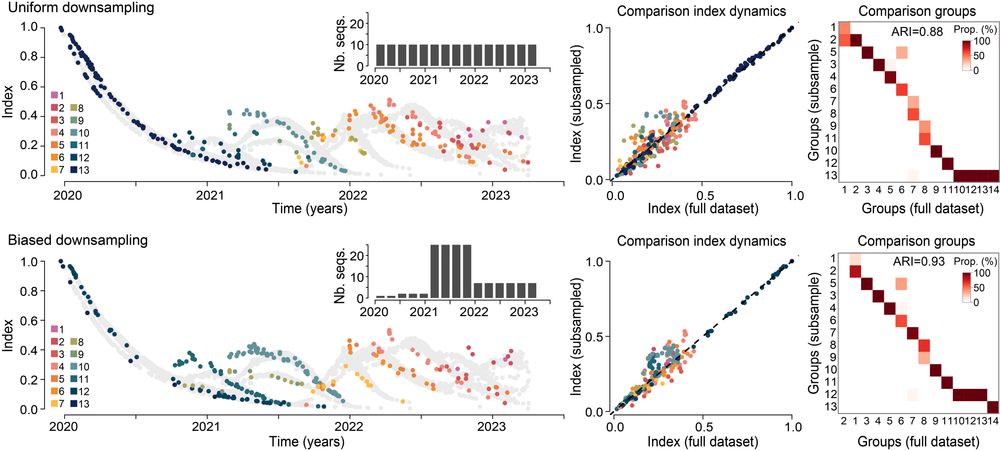
Importantly, phylowave is robust to uneven and limited observation. For example, we tested the framework on subsampled SARS-CoV-2 phylogenies of only 150 sequences and were still able to detect nearly all the lineages.
This makes phylowave applicable to a wide range of pathogens! 10/n
02.01.2025 08:04
👍 4
🔁 0
💬 1
📌 0
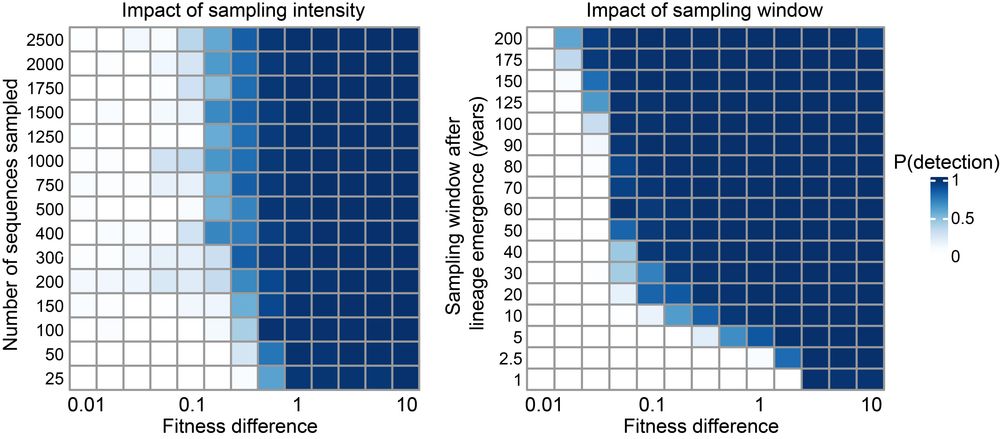
We tested phylowave using a simulation study. Its ability to recover lineages depends on the time between emergence and the dates of sequences, with lineages with only small fitness advantages requiring sequences covering longer time periods, as expected. 9/n
02.01.2025 08:04
👍 4
🔁 0
💬 1
📌 0
We show we can quantify the relative fitness of each lineage and identify genetic changes linked to the emergence of new, fitter lineages. This provides testable biological hypotheses about genetic variants in each pathogen that are driving the changes in population fitness of that pathogen. 8/n
02.01.2025 08:04
👍 4
🔁 1
💬 3
📌 0

We compared our lineage assignments with existing lineage definitions. We show we can recover the main known circulating lineages for each pathogen, as well as identify new lineages. For ex., phylowave detects three B. pertussis lineages with distinct index dynamics, not previously identified. 7/n
02.01.2025 08:04
👍 4
🔁 0
💬 1
📌 0

We applied phylowave to a broad set of viruses and bacteria: SARS-CoV-2, H3N2 influenza, Bordetella pertussis (bacterium causing whooping cough) and Mycobacterium tuberculosis. 6/n
02.01.2025 08:04
👍 3
🔁 0
💬 1
📌 0
We implemented a tree partitioning algorithm using generalised additive models that finds the set of lineages (i.e., groups of tips and nodes) that best explains the observed index dynamics. It allows for the automatic detection of circulating lineages based on differences in fitness. 5/n
02.01.2025 08:04
👍 2
🔁 0
💬 1
📌 0

The index is based on the expectation that nodes sampled from an emerging fitter lineage will be phylogenetically closer than the rest of the population at that time. Mathematically, we expect emerging fitter lineages to have different index dynamics than the rest of the population. 4/n
02.01.2025 08:04
👍 5
🔁 0
💬 1
📌 0
Phylowave enables to scan time-resolved phylogenies and sheds light on changes in population composition. It works by assigning an epidemic success index to each node (internal or terminal).
Here is an example on a SARS-CoV-2 phylogeny from nextstrain, coloured by nextstrain clades. 3/n
02.01.2025 08:04
👍 4
🔁 0
💬 1
📌 1
Existing methods to monitor the fitness of strains at the population level rely on pre-defined clade definitions, unlinked to underlying differences in fitness. We developed phylowave to tackle this issue and automatically detect lineages based on shared fitness and evolutionary relationships. 2/n
02.01.2025 08:03
👍 4
🔁 0
💬 1
📌 0
Learning the fitness dynamics of pathogens from phylogenies - Nature
Phylowave, an innovative phylogenetic approach, can identify the main circulating pathogen lineages with increased fitness and the associated genetic changes, enabling the timely identification of…
Incredibly excited to share that our manuscript was just published in @nature.com ! What a way to start the new year! 🎉
https://buff.ly/4gyYCzx
We present phylowave, a framework that enables to learn the fitness dynamics of pathogens from phylogenies.
🧵 A thread... 1/n
#IDSky #IDModelling
02.01.2025 08:03
👍 215
🔁 87
💬 3
📌 4
Congratulations Charlotte!
16.12.2024 13:41
👍 1
🔁 0
💬 0
📌 0

Graphic shows dramatic reduction in polio, rubella, mumps, measles, and hepatitis A cases after the introduction of vaccines for each disease.
Disease prevalence in US states before & after vaccine introduction
Graphic from Edward Tufte
More graphics.wsj.com/infectious-d... 🧪
15.11.2024 05:00
👍 3888
🔁 1700
💬 130
📌 166
Super resource to explore publicly available genome sequencing data! Currently supports H5N1, West Nile, RSV-A, RSV-B and SARS-CoV-2
GenSpectrum.org
#IDSky
25.11.2024 13:13
👍 6
🔁 0
💬 1
📌 0
Thrilled to have received this award, thank you @gensocuk.bsky.social! Many thanks to @hsalje.bsky.social @julianparkhill.bsky.social, Sylvain Brisse and the amazing research groups without whom the work wouldn't have been possible!
22.11.2024 13:23
👍 21
🔁 2
💬 2
📌 1

Infectious Disease Modelling #IDModelling
Join the conversation
Infectious Disease Modelling starter pack update! First pack is full so I created a second one. Pls keep on sending suggestions! (bio should contain experience relevant for this pack)
IDModelling pack 1: go.bsky.app/86Ao1a5
IDModelling pack 2 : go.bsky.app/2oBB7KX
22.11.2024 08:08
👍 53
🔁 26
💬 13
📌 7

Hey bluesky! It's great to see so many new folks jumping on the #IDSky wagon. If you're at #TropMed24 drop by my poster tomorrow at 11am. We can talk all things chikungunya burden, sex and age-dependent disease risk, and potential vaccine impact!
15.11.2024 15:00
👍 9
🔁 2
💬 0
📌 0
Infectious Disease Modelling starter pack!
Suggestions are welcome!
go.bsky.app/86Ao1a5
10.11.2024 18:06
👍 73
🔁 33
💬 13
📌 3















