
We have a little new paper at ICLR led by @AntonBushuiev.
Test time training for proteins :)
arxiv.org/abs/2411.02109
For example, you know the sequence that you want to fold, so fine-tune ESM on it at test time to get a better ESMFold structure prediction!
05.03.2026 15:56
👍 5
🔁 2
💬 0
📌 0
Joint work with @roman-bushuiev.bsky.social, and Olga Pimenova, Nikola Zadorozhny, Raman Samusevich, Elisabet Manaskova, @eunbelivable.bsky.social , @hannes-stark.bsky.social, Jiri Sedlar, @martinsteinegger.bsky.social , @pluskal-lab.org, @josef-sivic.bsky.social
05.03.2026 12:08
👍 3
🔁 0
💬 0
📌 0
More broadly, ProteinTTT can be applied to any protein language model to improve structure, fitness, or function prediction. In the new version of the paper we also extend ProteinTTT to autoregressive and discrete diffusion models, and add a deeper analysis of success and failure cases
05.03.2026 12:08
👍 0
🔁 0
💬 1
📌 0
If you have a protein where AlphaFold or other methods struggle (e.g., viral proteins, antibodies, or low-homology targets), give ProteinTTT a try. ProteinTTT adapts ESMFold to one protein at a time before predicting structure, boosting downstream accuracy for most proteins
05.03.2026 12:08
👍 3
🔁 1
💬 1
📌 0

ProteinTTT is now easy to run on Hugging Face Spaces and Google Colab. We’ll also be presenting the paper at ICLR 2026 🇧🇷
🤗 Hugging Face Space: huggingface.co/spaces/pimen...
⚙️ Google Colab: colab.research.google.com/drive/1l_h7c...
🧵👇
05.03.2026 12:08
👍 39
🔁 9
💬 3
📌 0

Mirdita Lab - Laboratory for Computational Biology & Molecular Machine Learning
Mirdita Lab builds scalable bioinformatics methods.
My time in @martinsteinegger.bsky.social's group is ending, but I’m staying in Korea to build a lab at Sungkyunkwan University School of Medicine. If you or someone you know is interested in molecular machine learning and open-source bioinformatics, please reach out. I am hiring!
mirdita.org
20.01.2026 11:07
👍 104
🔁 55
💬 7
📌 1
【論文簡単解説】One protein is all you need簡単解説💨 #shorts #vtuber準備中 #澪乃ゆい
YouTube video by Miono Yui ch. | 澪乃ゆい
Amazing summary of our recent ProteinTTT paper youtube.com/shorts/XWueh...
03.01.2026 13:25
👍 4
🔁 2
💬 1
📌 0
Very happy @roman-bushuiev.bsky.social and I joined the amazing team led by @hannes-stark.bsky.social to work on BoltzGen, a generative model for binder design based on Boltz-2. Excited what it will enable!
27.10.2025 12:34
👍 12
🔁 4
💬 0
📌 0

Excited to release BoltzGen which brings SOTA folding performance to binder design! The best part of this project is collaborating with a broad network of leading wetlabs that test BoltzGen at an unprecedented scale, showing success on many novel targets and pushing the model to its limits!
26.10.2025 22:40
👍 103
🔁 41
💬 3
📌 5
BIG BIG congratulations to our PhD student
@roman-bushuiev.bsky.social for receiving the Google PhD Fellowship 2025 in Health Research! 🎉💰 goo.gle/43wJWw8
25.10.2025 00:44
👍 21
🔁 3
💬 1
📌 1
Yes, exactly! This is an interesting perspective. Actually, running .ttt with msa_pth provided is effectively evotuning. We should highlight this connection better in the paper!
24.10.2025 02:53
👍 1
🔁 0
💬 0
📌 0
This is a joint work with @roman-bushuiev.bsky.social , and Olga Pimenova, Nikola Zadorozhny, Raman Samusevich, Elisabet Manaskova, @eunbelivable.bsky.social , @hannes-stark.bsky.social , Jiri Sedlar, @martinsteinegger.bsky.social , @pluskal-lab.org , @josef-sivic.bsky.social
23.10.2025 13:08
👍 5
🔁 0
💬 0
📌 0

Here, customization primarily benefits challenging, out-of-distribution proteins that are poorly represented in sequence databases (as measured by MSA size)
23.10.2025 13:08
👍 2
🔁 0
💬 1
📌 0

For example, ProteinTTT applied to ESMFold improves 19% of AlphaFold2-predicted viral protein structures in BFVD
23.10.2025 13:08
👍 3
🔁 0
💬 1
📌 0

This consistently improves performance of various protein language models across protein structure, fitness and function prediction, particularly on challenging targets
23.10.2025 13:08
👍 2
🔁 0
💬 1
📌 0

ProteinTTT overview
ProteinTTT enables *customizing* protein language models to one target protein at a time without assuming any additional data via on-the-fly self-supervised fine-tuning on the single protein
23.10.2025 13:08
👍 3
🔁 0
💬 1
📌 1
We train machine learning models on millions of proteins. But when it comes to making predictions, do we need them to understand all proteins at once? Often, we need an accurate model for the specific protein we are studying or designing. We address this with ProteinTTT arxiv.org/abs/2411.02109 1/🧵
23.10.2025 13:08
👍 68
🔁 25
💬 2
📌 0

⚛️ @pluskal-lab.org from IOCB Prague, together with his student @roman-bushuiev.bsky.social and colleagues from #CIIRC CTU, Josef Šivic and @anton-bushuiev.bsky.social, have developed a machine learning model called #DreaMS – which accelerates the analysis of previously unknown molecules.
28.05.2025 13:42
👍 12
🔁 5
💬 1
📌 0
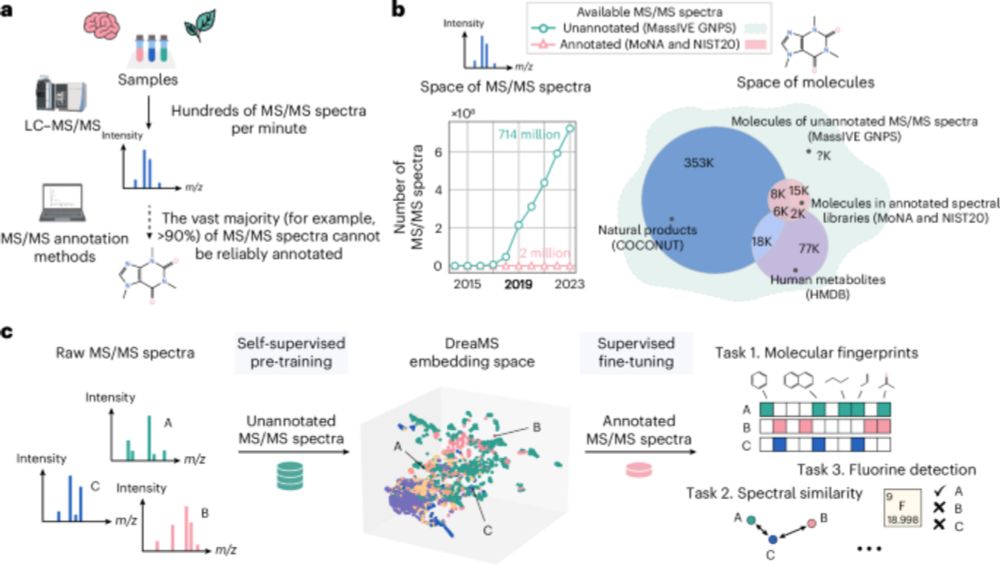
Self-supervised learning of molecular representations from millions of tandem mass spectra using DreaMS - Nature Biotechnology
A transformer model is used to construct the DreaMS Atlas—a molecular network of 201 million MS/MS spectra.
Mass spectrometry is a key method to discover and identify molecules in biological and environmental samples. Yet, >90% of mass spectra remain hard to interpret. In our recent paper, we present DreaMS — a foundation model to interpret mass spectra of small molecules.
www.nature.com/articles/s41...
26.05.2025 13:44
👍 25
🔁 10
💬 1
📌 0
This paper represents a great effort by @roman-bushuiev.bsky.social and his brother @anton-bushuiev.bsky.social. The DreaMS foundation model for mass spectra of small molecules now opens lots of avenues for possible downstream applications. It might be a game changer for computational metabolomics.
24.05.2025 08:21
👍 52
🔁 20
💬 0
📌 1

Back in July 2023 we organized a small bioML symposium at @iocbprague.bsky.social, and it turned out to be a very pleasant and successful event. This summer we are following up with a great line-up of speakers. Please register for free, deadline May 30. tinyurl.com/bioMLPrague25
11.04.2025 11:34
👍 17
🔁 8
💬 0
📌 0

🚀 Exciting MassSpecGym leaderboard update 🚀
Two new machine learning models achieve up to a 300% improvement in de novo molecular generation given mass spectra and corresponding chemical formulae. 🔥 1/n
07.03.2025 14:58
👍 11
🔁 3
💬 1
📌 0
MassSpecGym - the first comprehensive benchmark for the discovery and identification of molecules from MS/MS data.
@roman-bushuiev.bsky.social
@anton-bushuiev.bsky.social
@josef-sivic.bsky.social
@pluskal-lab.org
NeurIPS 2024 paper: arxiv.org/abs/2410.23326
#ChemSky #MassSpec #AI4Science
20.02.2025 17:16
👍 8
🔁 4
💬 1
📌 0

🤝 In April 2024, brothers Roman and Anton Bushuiev from the teams of @pluskal-lab.org @iocbprague.bsky.social and Josef Šivic #CIIRC_CTU initiated a collaboration among 14 research institutes across the globe to benchmark #AI methods for the discovery of molecules from mass spectrometry data. 1/2
20.02.2025 14:06
👍 11
🔁 5
💬 1
📌 1
MassSpecGym: A benchmark for the discovery and identification of molecules
YouTube video by PolarisHQ
MassSpecGym is the largest publicly available collection of mass spectra data with 231K spectra for 29K unique molecular structures. 33% of the dataset was generated from newly measured, in-house data.
🛡️The dataset is now certified on Polaris! polarishub.io/datasets/rom...
youtu.be/G8ZnVRm0ogc
24.01.2025 18:55
👍 16
🔁 5
💬 1
📌 0
The MassSpecGym team presenting the poster at #NeurIPS2024! Fei Wang,
Adamo Young,
@roman-bushuiev.bsky.social,
@anton-bushuiev.bsky.social,
Raman Samusevich
14.12.2024 17:27
👍 20
🔁 3
💬 0
📌 0

Met the twins @roman-bushuiev.bsky.social @anton-bushuiev.bsky.social
13.12.2024 01:43
👍 14
🔁 2
💬 0
📌 0



















