
Figure 1. Cryo-EM of abundant protein complexes in native membranes.

Figure 2. Cryo-EM of membrane proteins in vesicles.

Figure 3. 3D reconstruction of V-ATPase in native synaptic vesicle membranes.

Figure 4. Generation of membrane vesicles for structure determination of proteins in their native lipid bilayer.
I've written a review on what I think is an extremely exciting direction in cryo-EM:
Cryo-EM of endogenous membrane proteins in their native lipid bilayer
Open access in Quarterly Reviews of Biophysics:
doi.org/10.1017/S003...
06.03.2026 12:40
👍 152
🔁 56
💬 3
📌 1

Structure of the ribosome-bound SND3 translocon complex from the heat-tolerant fungus Chaetomium thermophilum. The SND3 translocon inserts proteins into the endoplasmic reticulum membrane as they are produced by the ribosome.
SNDing proteins into the membrane! Our new publication from @melaniemcdowell.bsky.social ’s group identifies the SND3 protein as a new route for membrane protein insertion! 🍄 📘 Read more here: www.mpg.de/25599408/102... Image: Louise Duever.
30.10.2025 08:55
👍 15
🔁 8
💬 2
📌 1
Postdoctoral researcher in structural bioenergetics (m/w/d)
Prof. Hartmut Michel's group @mpibp.bsky.social is recruiting a postdoc in structural bioenergetics. See the link for more detail.👇
24.10.2025 09:36
👍 2
🔁 1
💬 0
📌 0
This one is a bit of a departure from the usual and definitely a work in progress!
We found that by using ab initio reconstruction at very high res, in very small steps, we could crack some small structures that had eluded us - e.g. 39kDa iPKAc (EMPIAR-10252), below.
Read on for details... 1/x
13.09.2025 00:04
👍 266
🔁 82
💬 20
📌 14
That's a wrap! The results of the first #cryoEM heterogeneity challenge are up on biorxiv!
biorxiv.org/content/10.110
23.07.2025 21:43
👍 45
🔁 22
💬 3
📌 4
Proud to share our first lab pre-print: “SND3 is the membrane insertase within a fungal multipass translocon” where @tzujingyang.bsky.social solved the structure of a ribosome-associated SND3-translocon complex involved in ER membrane protein insertion ➡️ doi.org/10.1101/2025...
12.07.2025 00:49
👍 92
🔁 32
💬 3
📌 4
Congrats 🎉
15.06.2025 10:05
👍 1
🔁 0
💬 1
📌 0
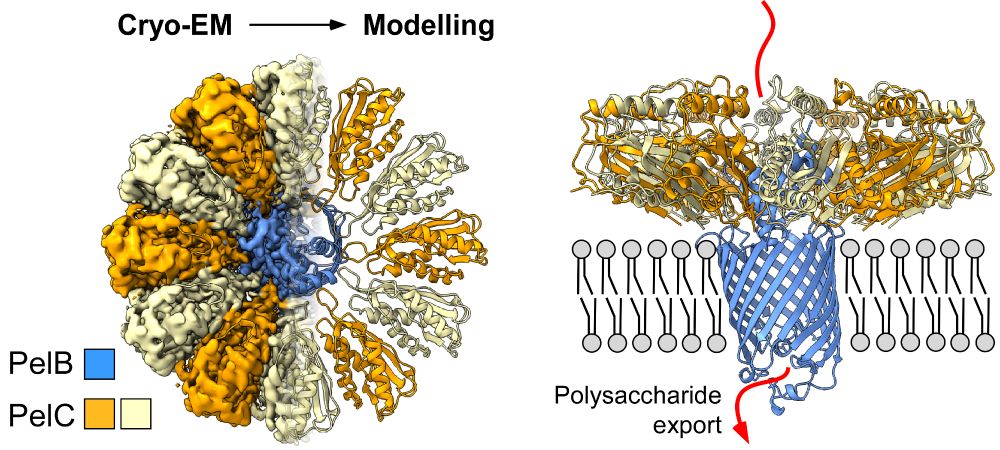
It is online since a few days, so time to recap: Our first steps in a new field of bacterial polysaccharide secretion, focused on Pel system from Pseudomonas aeruginosa:
www.nature.com/articles/s41...
15.06.2025 08:41
👍 50
🔁 15
💬 4
📌 1

Substrate-induced assembly and functional mechanism of the bacterial membrane protein insertase SecYEG-YidC
The universally conserved Sec translocon and the YidC/Oxa1-type insertases mediate biogenesis of alpha-helical membrane proteins, but the molecular basis of their cooperation has remained disputed over decades. A recent discovery of a multi-subunit insertase in eukaryotes has raised the question about the architecture of the putative bacterial ortholog SecYEG-YidC and its functional mechanism. Here, we combine cryogenic electron microscopy with cell-free protein synthesis in nanodiscs to visualize biogenesis of the polytopic membrane protein NuoK, the subunit K of NADH-quinone oxidoreductase, that requires both SecYEG and YidC for insertion. We demonstrate that YidC is recruited to the back of the translocon at the late stage of the substrate insertion, in resemblance to the eukaryotic system, and in vivo experiments indicate that the complex assembly is vital for the cells. The nascent chain does not utilize the lateral gate of SecYEG, but enters the lipid membrane at the SecYE-YidC interface, with YidC being the primary insertase. SecYEG-YidC complex promotes folding of the nascent helices at the interface prior their insertion, so the examined cellular pathway follows the fundamental thermodynamic principles of membrane protein folding. Our data provide the first detailed insight on the elusive insertase machinery in the physiologically relevant environment, highlight the importance of the nascent chain for its assembly, and prove the evolutionary conservation of the gate-independent insertion route. ### Competing Interest Statement The authors have declared no competing interest. Deutsche Forschungsgemeinschaft, https://ror.org/018mejw64, Ke1879/3, 267205415 (CRC 1208) European Research Council, https://ror.org/0472cxd90, CRYOTRANSLATION
Very special feelings to announce this one... A project that started like 10 years ago is reaching the finish line, ready to shine. In a dream-team with @beckmannlab.bsky.social we solved the long-chased structure of the active membrane protein insertase SecYEG-YidC
www.biorxiv.org/content/10.1...
27.05.2025 09:21
👍 60
🔁 24
💬 7
📌 7
Beautiful work! Congrats 🎉
28.05.2025 21:30
👍 2
🔁 0
💬 0
📌 0
We are happy to share that our snR30 story is finally out in @natureportfolio.nature.com 🥳 We report the first structure of a H/ACA snoRNP acting in ribosome synthesis thereby providing a detailed structural and biochemical view of the snR30 snoRNP guiding local 18S rRNA subdomain folding. 👇👇👇
26.05.2025 09:29
👍 62
🔁 27
💬 1
📌 3
New preprint on 3D heterochromatin architecture in human cells! Great collab with @sergiocruzleon.bsky.social & @johannesbetz.bsky.social from @hummerlab.bsky.social, @marinalusic.bsky.social & the Turoňová lab. Many thanks to my supervisor @becklab.bsky.social. bioRxiv: tinyurl.com/3a74uanv 🧵👇
11.04.2025 09:03
👍 224
🔁 67
💬 3
📌 4

Towards community-driven visual proteomics! Excited to finally share this large-scale curated & annotated dataset of 1829 high-quality #cryoET tomograms of the little green alga that just keeps giving— Chlamydomonas! 🧪🧶🧬🌾🌊🌍
Preprint📜: www.biorxiv.org/content/10.1...
A short thread🧵👇
06.01.2025 11:18
👍 339
🔁 129
💬 7
📌 16

Chaperonin TRiC within human cells. The left panel displays a representative 3D map from a tomogram illustrating the spatial distribution of open and closed TRiC complexes in human cells. The top right panel highlights the interaction of PDCD5 with the CCT3-1-4 subunits of the open TRiC complex. The bottom right panel shows the closed TRiC complex forming clusters of two
particles.
Do not miss our latest #cryoET work! 🕵️ Xing et al. present the dynamic architecture of TRiC in human cells! The study of this protein-folding chaperonin opens the door to understanding its dysfunction in diseases like cancer and neurodegeneration 🧠🧪 #teamTOMO @becklab.bsky.social
12.12.2024 09:16
👍 28
🔁 9
💬 1
📌 0
I see. So probably the first frame affects more on a clean, homogeneous particle stack? And yes there's usually ~0.2 Å improvement after polishing for the high-res structures.
29.11.2024 14:26
👍 0
🔁 0
💬 0
📌 0
Interesting. With 1 e-/Å2/fm we constantly get 2-3.5 Å res (depending on samples) without discarding the first frame using MotionCor2. In your experience, how much improvement can benifit from discarding the first frame? Or is this only for MotionCor3?
28.11.2024 18:29
👍 0
🔁 0
💬 1
📌 0
Faster and more accurate protein modeling with CryFold. By focusing on nearby points (Local Attention) and employing map masking, CryFold excels in generating models from challenging maps regions. It enables a more cost-efficient model building. 🧵
💾 github.com/SBQ-1999/Cry...
25.11.2024 17:24
👍 111
🔁 38
💬 1
📌 3
Ok, trying to bring my community back together... I created a starter pack for lipids and membrane protein aficionados!
Feel free to share, add and distribute, so we can come together again! 😍
go.bsky.app/AEeXh86
16.11.2024 03:53
👍 124
🔁 65
💬 71
📌 7













