🎉 New year, NEW PREPRINT!
Bacteria exhibit astonishing genetic diversity, but where do new genes come from?
My best friend Arya Kaul (/labmate in the @baym lab) investigates how advantageous deletions can spawn new genes - "deletion-born fusions." 🧵:
06.01.2026 16:09
👍 49
🔁 30
💬 1
📌 2

What is the best strategy to win any contest?
Eliminate your opponents of course.
Recently, my friend @fernpizza.bsky.social showed how plasmids compete intracellularly (check out his paper published in Science today!). With @baym.lol, we now know they can fight.
www.biorxiv.org/content/10.1...
20.11.2025 22:11
👍 79
🔁 42
💬 3
📌 6
hey bluesky 👋 visa hurdles mean I’m looking for opportunities outside the US. I’m a computational biologist (bacterial + phage genomics, postdoc in Koonin’s group @ NIH). I am interested in teaming up on funding apps. reach out if this resonates!
15.09.2025 17:26
👍 69
🔁 89
💬 1
📌 3

ESGMAP is now on LinkedIn and Bluesky!
We published our first newsletter (lnkd.in/dUe43t8y). It introduces us, our mission, activities, and ways to get involved. Fill out our members' survey, and associate your publications with the group.
Follow us for updates on the mobile elements and plasmids.
19.06.2025 08:56
👍 5
🔁 4
💬 0
📌 1

The Wadsworth Center is expanding it's research focus on #virology /vector-borne diseases!
www.asmcareerconnections.org/job/research-scientist-5-g-31/78488664/
This is a unicorn assistant prof equivalent position with 12-month hard money salary and no teaching commitments (!)
Reposts appreciated 🙏
03.06.2025 21:35
👍 49
🔁 60
💬 3
📌 4
Job Details
New Research Assistant position available in my lab to work on a project testing the efficacy of pills dependent #phage against clinical isolates of enteric bacteria . Part of a growing phage research theme in my lab my.corehr.com/pls/uoxrecru...
08.05.2025 21:24
👍 28
🔁 26
💬 2
📌 1
The link is not working for me 😔
25.04.2025 09:58
👍 1
🔁 0
💬 1
📌 0
Any idea when the abstract submission portal opens for IMMEM XIV? 😊 @esgem-sg.bsky.social
03.03.2025 09:12
👍 0
🔁 0
💬 0
📌 0
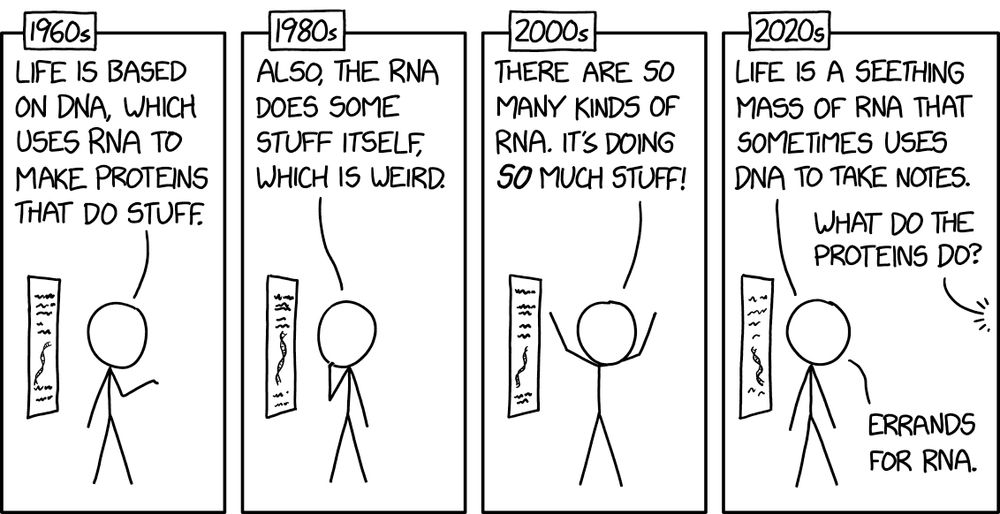

RNA xkcd.com/3056
26.02.2025 14:58
👍 17849
🔁 2559
💬 154
📌 171
Ground-breaking work by one of the most amazing of scientists. A must read for all plasmids afficionado (and and any evolutionary biologist in general!)
21.02.2025 22:17
👍 8
🔁 3
💬 0
📌 0
It seems really good but apparently rough around the edges regarding the UI, so I am waiting a bit for them to patch it first!
06.02.2025 17:30
👍 1
🔁 0
💬 1
📌 0
This paper has been a long time coming: We looked at the genomes of historical bacterial samples over a century to look for trends of antibiotic resistance genes, finding multiple instances of them in infections before the age of antibiotics, but an increase in both frequency and mobility after
17.01.2025 20:53
👍 305
🔁 122
💬 11
📌 5
Genomic resistance in historical clinical isolates increased in frequency and mobility after the age of antibiotics https://www.biorxiv.org/content/10.1101/2025.01.16.633422v1
17.01.2025 16:17
👍 17
🔁 4
💬 2
📌 2
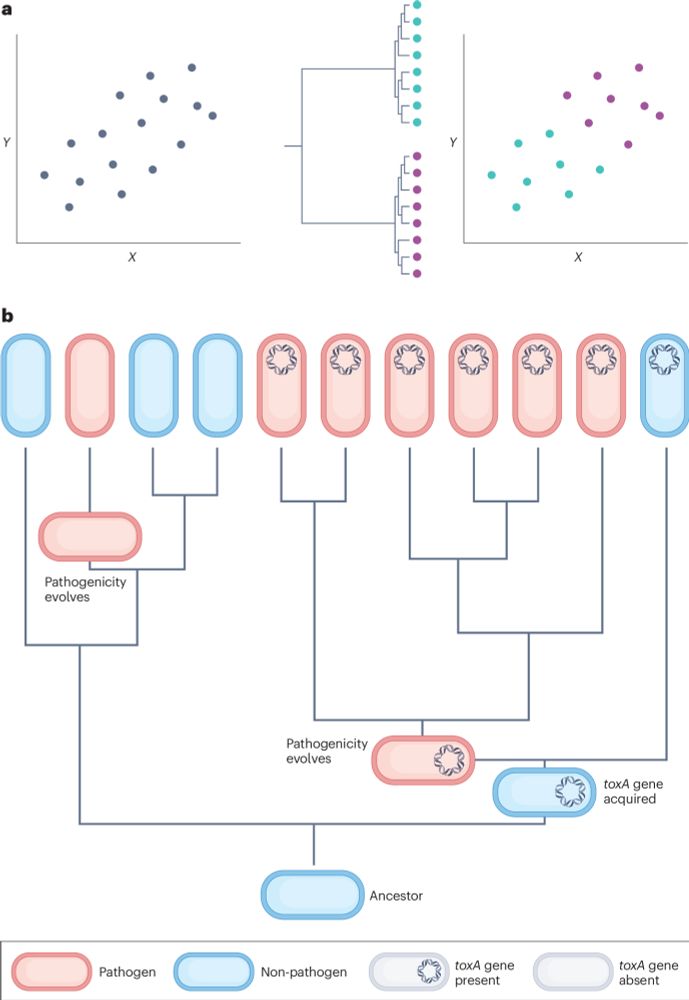
A phylogenetic approach to comparative genomics
Nature Reviews Genetics - Controlling for phylogeny is essential in comparative genomics studies, because species, genomes and genes are not independent data points within statistical tests. The...
Our review is out in Nature Reviews Genetics! rdcu.be/d5AY2
We show how phylogeny-based methods can resolve the problem of non-independence in genomic datasets.
These methods must be considered an essential part of the comparative genomics toolkit.
@lauriebelch.bsky.social @stuwest.bsky.social
08.01.2025 13:19
👍 194
🔁 95
💬 5
📌 4
Sharing three fully funded PhD opportunities to join my lab at Queen’s to work on topics spanning bacterial pathogens, antibiotic resistance, microbial interactions, & mobile genetic elements 🦠. Closing dates in January and February, and open to international candidates 🌍 Projects detailed below ⬇️
30.12.2024 14:26
👍 37
🔁 47
💬 4
📌 3
Gosh parts of me hate the concept of ChatGPT with a passion, but I have to acknowledge it's supremacy when trying to debug a Snakemake pipeline, which I am not convinced are made to be understood by humans 😅
03.12.2024 10:47
👍 3
🔁 0
💬 0
📌 0
Volume 2 of the #AMR Starter Pack is here!
go.bsky.app/EUY2FUJ
Follow these amazing people along with those in Vol. 1
go.bsky.app/KkZfvrh
And don't forget to pin the Antimicrobial Resistance feed to get all the AMR posts in one place
bsky.app/profile/did:...
02.12.2024 06:52
👍 36
🔁 22
💬 9
📌 7
Don't know how to do polls in Bluesky, but am very curious to know how what the #microsky thinks.
Genes that increase fitness in natural conditions are:
A. Over-represented on plasmids
B. Over-represented on the chromosome
C. Evenly distributed between plasmids and the chromosome
27.11.2024 21:11
👍 1
🔁 1
💬 5
📌 0
BRIG
For circular plots, I have found that out of the box elderly BRIG beatsonlab.com/softwares/br... remains the best. Otherwise for more custom plots I usually go with circos-like packages (like pyCirclize in Python) but it is much more work
01.12.2024 18:36
👍 2
🔁 0
💬 0
📌 0
That's low-key one of my favorite thing about Oslo - the city center is very often car free, and the few cars are often electric, which makes them much less audible.
Going to back to a 'proper' big city (Stockholm) for a week felt weird!
27.11.2024 08:59
👍 10
🔁 0
💬 0
📌 0
Please sign up, it will be fun!
Open for all with an interest in AMR and genomics (in the UK and elsewhere).
www.targetamr.org.uk
(and please give @target-amr.bsky.social a follow!)
30.10.2024 11:56
👍 22
🔁 23
💬 5
📌 0
Signed up! 😊
24.11.2024 14:35
👍 1
🔁 0
💬 0
📌 0