
I am incredibly excited to share that I will start my independent lab at the @unidue-zmb.bsky.social at the @unidue.bsky.social as Junior Professor of Cellular Biochemistry. Research in my lab has the goal to decipher the ubiquitin code!
There are multiple open positions!
(1/3)
05.03.2026 11:00
👍 79
🔁 20
💬 11
📌 4
CryoEM data collected at the ever helpful @scmicryoem.bsky.social
27.02.2026 08:50
👍 2
🔁 0
💬 1
📌 0

Comparison of PCNA and FANCD2
We call the SLiM found on these proteins a FANCD2-interacting protein box (DIP-box), as a shout-out to PCNA-interacting protein boxes (PIP-boxes)
27.02.2026 08:46
👍 2
🔁 0
💬 1
📌 0
Combining information from these structures with functional information from #uniprot and co-evolutionary information via #alphafold we found new FANCD2 interactors that utilize the same site
27.02.2026 08:45
👍 5
🔁 3
💬 1
📌 0

CryoEM consensus reconstructions of FANCI-FANCD2ub with FAN1 or CtIP
While figuring out the structural basis of FAN1 and CtIP interactions with the FANCD2 DNA clamp, we noticed a common interaction point that USP1 also uses
27.02.2026 08:41
👍 1
🔁 0
💬 1
📌 1
Excited to share our new paper uncovering how FANCD2 recognises various DNA repair proteins www.biorxiv.org/content/10.6... @hw1o9.bsky.social
27.02.2026 08:41
👍 15
🔁 8
💬 1
📌 1

We posted a biorxiv preprint on structural bioinformatics, AlphaFold modeling & machine learning on predicting specificity of E3 ligase ring domains for different E2 enzymes. 1/4
Preprint: www.biorxiv.org/content/10.6...
Models/data (UbiqCore website): dunbrack.fccc.edu/ubiqcore
17.02.2026 05:02
👍 52
🔁 27
💬 1
📌 3

AI in Protein Design: Hype vs. Reality Explained by David Baker
In this GEN interview, Nobel Laureate David Baker, PhD, emphasizes that designing proteins from scratch is reality and unpacks what's needed for AI to transform medicine.
AI in Protein Design: Hype vs. Reality Explained by David Baker
In this GEN interview, he emphasizes that designing new proteins from scratch is now a reality. Whether AI transforms medicine will require improving our understanding of biology's complexity
02.11.2025 17:34
👍 13
🔁 8
💬 0
📌 2
I love AlphaFold—but please include PAE (Predicted Aligned Error) plots for every “interaction.” Pretty PDBs ≠ proof. If the PAE doesn’t show an interface, it ain’t one. Show the plots. Structural biology's reputation is on the line.
24.10.2025 17:44
👍 129
🔁 29
💬 5
📌 4
Where next for structural bioinformatics?
I wrote a blog post about the future of structural bioinformatics.
Where to go after AlphaFold? How do we avoid the field becoming a load of half-baked LLMs?
Let me know what you think.
jgreener64.github.io/posts/struct...
15.10.2025 14:16
👍 39
🔁 14
💬 4
📌 2
Will Reform win the next UK election?
YouTube video by Garys Economics
Will Reform win the next UK election?
www.youtube.com/watch?v=mcu0...
21.09.2025 08:38
👍 110
🔁 24
💬 9
📌 7
RFdiffusion3 is here! We train a general network that explicitly models every atom and use it to design active enzymes and DNA binders.
Tremendous team effort with Jasper, Rohith, Raktim, Rafi, Yanjing, Paul, Jonathan, and many others!
Check it out: lnkd.in/eiUFfJaM.
19.09.2025 15:45
👍 31
🔁 9
💬 1
📌 0
This one is a bit of a departure from the usual and definitely a work in progress!
We found that by using ab initio reconstruction at very high res, in very small steps, we could crack some small structures that had eluded us - e.g. 39kDa iPKAc (EMPIAR-10252), below.
Read on for details... 1/x
13.09.2025 00:04
👍 266
🔁 82
💬 20
📌 14
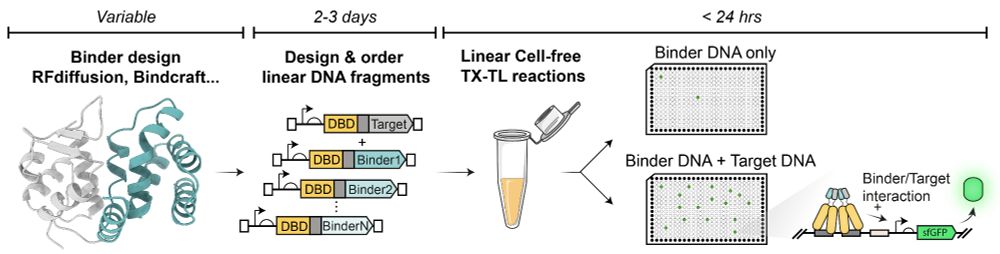
You designed binders for your favourite protein and wish there was a way to experimentally screen them within 24h w/ only a set of pipettes and a plate reader?
Check out our Cell-Free 2-Hybrid approach (CF2H)
Full post: tinyurl.com/48cz5nb6
Preprint: www.biorxiv.org/cgi/content/...
17.07.2025 11:41
👍 67
🔁 23
💬 0
📌 3
I'm back – and I think we can win
YouTube video by Garys Economics
I'm back. Here's how we fix the UK's dire economic and political situation www.youtube.com/watch?v=LMor...
13.07.2025 08:11
👍 375
🔁 111
💬 27
📌 19
Folddisco finds similar (dis)continuous 3D motifs in large protein structure databases. Its efficient index enables fast uncharacterized active site annotation, protein conformational state analysis and PPI interface comparison. 1/9🧶🧬
📄 www.biorxiv.org/content/10.1...
🌐 search.foldseek.com/folddisco
07.07.2025 08:21
👍 153
🔁 70
💬 8
📌 3


This is quite something - 68% success rate for binder design w/ Chai 2, with many in the low-nM to pM range - impressive stuff chaiassets.com/chai-2/paper...
02.07.2025 15:19
👍 16
🔁 2
💬 0
📌 0

We have written up a tutorial on how to run BindCraft, how to prepare your input PDB, how to select hotspots, and various other tips and tricks to get the most out of binder design!
github.com/martinpacesa...
30.06.2025 19:45
👍 138
🔁 55
💬 4
📌 0

The E3 ligase Parkin ubiquitinates many proteins. Koszela, Walden (@hw1o9.bsky.social) et al. @uofglasgow.bsky.social identify and biochemically validate a direct Parkin interaction with a substrate, Miro1. rupress.org/jcb/article/...
27.06.2025 20:10
👍 36
🔁 13
💬 0
📌 3
A nifty trick our lab is using to improve structure prediction of viral membrane proteins in #AlphaFold3! 👇
18.06.2025 13:43
👍 29
🔁 8
💬 2
📌 0

🧬 New from crystal structures from our lab in EMBO Journal! We reveal how the FANCM enzyme, an emerging therapeutic target it cancer, has evolved to specifically recognise branched DNA and activate the Fanconi anaemia pathway of DNA repair. doi.org/10.1038/s443... 🧵👇
02.06.2025 00:41
👍 30
🔁 5
💬 2
📌 0
We are happy to release LocScale2.0: a tool for context-aware, confidence-weighted cryoEM map optimisation.
Taking two half maps as input, LcoScale-2.0 produces feature-enhanced maps along with a robust confidence score that guides objective map interpretation.
cryotud.github.io/locscale/
22.05.2025 10:10
👍 121
🔁 38
💬 1
📌 2

New from #JRSocInterface: Emerging frontiers in protein structure prediction following the #AlphaFold revolution. Read more: royalsocietypublishing.org/doi/10.1098/...
21.04.2025 16:13
👍 2
🔁 3
💬 1
📌 0
We also cover confidence metrics and suggest some guidelines for publishing bespoke AlphaFold predictions.
22.04.2025 16:46
👍 1
🔁 0
💬 0
📌 0

Different classes of protein structure prediction sorted by geometric complexity and evolutionary information
We speculate on the difficulty of developing prediction tools that are generally applicable to each prediction class by breaking down into geometric complexity (e.g. number of possible bond types) and evolutionary information (e.g. embedded in MSAs).
22.04.2025 16:46
👍 0
🔁 0
💬 1
📌 0
Our review on #AlphaFold focussing on prediction classes beyond simple protein monomers (PPIs, conformational changes, protein-ligand complexes, etc) royalsocietypublishing.org/doi/10.1098/...
22.04.2025 16:46
👍 8
🔁 3
💬 1
📌 0
Excited to share our preprint “BoltzDesign1: Inverting All-Atom Structure Prediction Model for Generalized Biomolecular Binder Design” — a collaboration with
@martinpacesa.bsky.social, @Zhidian Zhang, @Bruno E. Correia, and @sokrypton.org
🧬 Code will be released in a couple weeks
08.04.2025 12:17
👍 62
🔁 17
💬 5
📌 1





















