
A pictographic model of chromatin folding in the nucleus. DNA wrapped around nucleosomes aggregates into nanoscale domains, whilst newly nucleosome depleted DNA that forms after acute degradation of FACT projects from these domains and engages in long-range interactions that can span TAD boundaries
New preprint from @anadopico.bsky.social (who I have had the great pleasure of supervising with Tom Milne) and our fantastic collaborators in the @davieslab.bsky.social
www.biorxiv.org/content/10.6...
#genomics #GeneRegulation #chromatin
19.02.2026 16:10
👍 13
🔁 8
💬 1
📌 0
Excited to share that a part of my PhD work is online now on BioRxiv!
www.biorxiv.org/content/10.6...
We developed a genome editing approach to target the JAK2 V617F mutation and demonstrate its potential to alleviate MPN hallmarks in primary patient cells.
Thread 👇
20.12.2025 09:36
👍 22
🔁 8
💬 4
📌 0
Our Science paper is out!
Huge congratulations to @huabin-zhou.bsky.social, Mike Rosen, and the brilliant @janhuemar.bsky.social @juliamaristany.bsky.social and @kieran-russell.bsky.social from our group
News: bit.ly/4avnkAr and bit.ly/3XBGVHS
Great perspective by @vram142.bsky.social +K Zhang
05.12.2025 09:47
👍 87
🔁 41
💬 2
📌 1
Fascinating paper on a number of levels…
Anyone else think that NASA are basically saying that it’s likely that DNA/RNA/protein based life is likely to predate the solar system and arrived on earth on a comet…
So there are likely to be DNA based life forms throughout the universe?
02.12.2025 20:22
👍 2
🔁 0
💬 0
📌 0
Thanks so much for highlighting our images created with @rcollepardo.bsky.social
01.12.2025 16:01
👍 8
🔁 2
💬 0
📌 0

Oxford scientists capture genome’s structure in unprecedented detail
RDM scientists have achieved the most detailed view yet of how DNA folds and functions inside living cells, revealing the physical structures that control when and how genes are switched on.
Scientists have the most detailed view yet of how DNA folds and functions inside living cells!
The breakthrough helps us understand how genetic differences lead to disease and opens up fresh routes for drug discovery 👇
shorturl.at/nQjsx
@jojdavies.bsky.social @imm.ox.ac.uk @medsci.ox.ac.uk
06.11.2025 11:18
👍 4
🔁 1
💬 0
📌 1

Happy to share our latest publication, in which we show that the arrangement of nucleosomes around CTCF sites contributes to higher-order organisation of chromatin into TADs: www.embopress.org/doi/full/10....
27.10.2025 11:49
👍 71
🔁 31
💬 0
📌 2
Thanks Elzo! It has been a real pleasure collaborating with you on this.
05.11.2025 19:50
👍 1
🔁 0
💬 0
📌 0
Yes that’s correct - they are due to ligation junctions between the linkers of adjacent nucleosomes which have a fixed distance between them but a variable position with respect to the DNA sequence.
05.11.2025 18:58
👍 0
🔁 0
💬 1
📌 0


Not from Tron or a psychedelic wallpaper. This exquisite pic reveals chromatin at base-pair resolution, captured by #ListerFellow James Davies and collaborators🤩
"For the first time, we can see how the genome's control switches are physically arranged inside cells." @jojdavies.bsky.social
05.11.2025 16:50
👍 12
🔁 6
💬 1
📌 0

Finally, we propose a model in which chromatin is self-organising based on histone acetylation and nucleosome depletion. We hypothesise that active cis-regulatory elements may contact one another at or above a layer of acetylated nucleosomes.
05.11.2025 17:17
👍 18
🔁 2
💬 1
📌 0
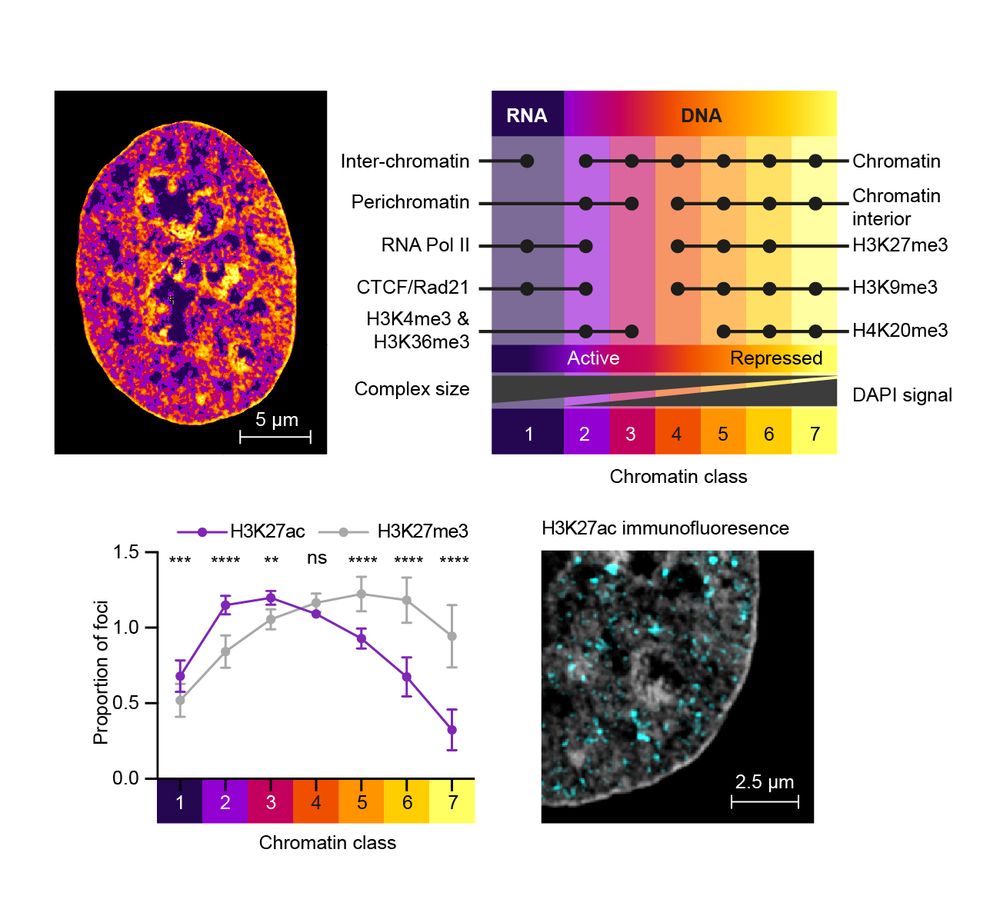
We then collaborated with Lothar Schermelleh to undertake super-resolution microscopy of chromatin acetylation. In keeping with his previous work we find that histone acetylation maps to regions of chromatin that are adjacent to the interchromatin compartment.
05.11.2025 17:17
👍 3
🔁 0
💬 1
📌 0


And histone acetylation drive the formation of the nanoscale domains we see in the MCC data.
This leads us to conclude that the biophysical properties of chromatin, particularly nucleosome depletion and histone acetylation, drive fine scale structure.
05.11.2025 17:17
👍 1
🔁 0
💬 1
📌 0

We show that nucleosome depletion....
05.11.2025 17:17
👍 0
🔁 0
💬 1
📌 0
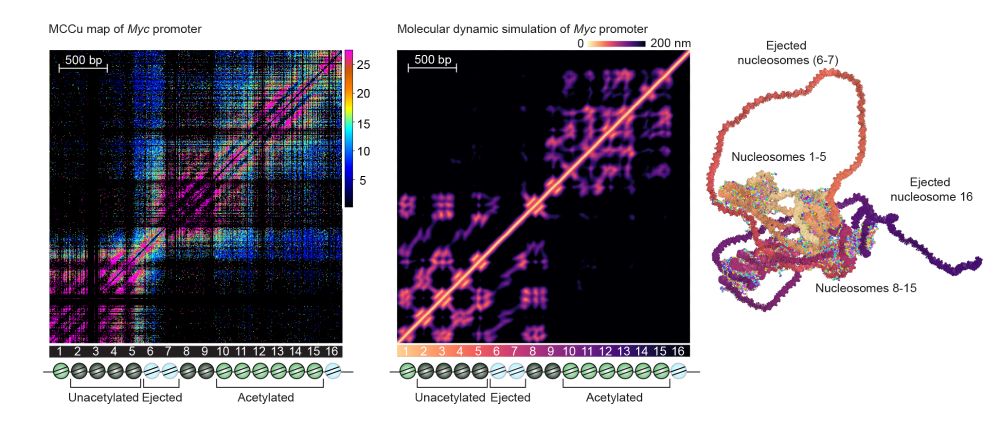
In collaboration with Rosana Collepardo-Guevara who has develops a chemically specific molecular dynamic simulation of the physicochemical properties of chromatin, we constructed an MD simulation of the Myc promoter that closely matches the MCC data.
05.11.2025 17:17
👍 2
🔁 0
💬 1
📌 0

However, at other sites only the intricate structure of the nucleosome depleted region changes.
From this we conclude that nucleosome depletion itself is a major driver of long-range contacts.
05.11.2025 17:17
👍 3
🔁 1
💬 1
📌 0

At some sites SOX2 depletion causes complete collapse of the nucleosome depleted region at the enhancer and loss of all contacts.
05.11.2025 17:17
👍 3
🔁 0
💬 1
📌 0
In collaboration with Rob Klose and Elzo de Wit, we explored the role of depletion of the mediator complex components (MED14, MED13 and MED1) and transcription factors (SOX2 and NANOG).
05.11.2025 17:17
👍 0
🔁 0
💬 1
📌 0

We are able to map specific structures within the nucleosome depleted regions at regulatory elements that are driven by transcription factors.
05.11.2025 17:17
👍 2
🔁 0
💬 1
📌 0

Between nucleosome depleted regions we see nanoscale domains, that appear similar to TADs but are approximately 100- to 1000-fold smaller
05.11.2025 17:17
👍 4
🔁 0
💬 2
📌 0

Our latest paper has just been published in Cell!
doi.org/10.1016/j.ce...
We developed a new method called MCC ultra, which allows 3D chromatin structure to be visualised with a 1 base pair pixel size.
05.11.2025 17:17
👍 209
🔁 79
💬 6
📌 11

We’re really excited to see Hangpeng Li present our latest work at the CSH Mechanisms of Eukaryotic Transcription meeting.
27.08.2025 20:48
👍 5
🔁 3
💬 0
📌 0
Thrilled that our paper is in print @science.org!!
*Platelets sequester cell free DNA, including free fetal and tumour-derived DNA*
Tweetorial from @l-cmurphy.bsky.social below. Check out the news feature science.org/content/arti... and terrific editorial from Dennis Lo #platelets_in_the_limelight
15.08.2025 05:37
👍 73
🔁 22
💬 4
📌 3






















