
Excited to purchase the pipette sets and sample coolers—tools I used during my time in the Suvà lab.
Thank you, Mitsubishi Foundation, for supporting our research!

Excited to purchase the pipette sets and sample coolers—tools I used during my time in the Suvà lab.
Thank you, Mitsubishi Foundation, for supporting our research!
14/ Big thanks also to the institutions that provided tumor samples @dukepress.bsky.social, @mdanderson.bsky.social, Tokyo University Hospital, Pitié-Salpêtrière Hospital, St. Michael's Hospital, Seoul National University and NORLUX and funders NIH, NCI, Mark Foundation and Sontag Foundation.
13/ @weizmanninstitute.bsky.social, Mass General Brigham (Cancer Center, Pathology), @broadinstitute.org, @yalepress.bsky.social and University of Miami.
12/ Big thanks to the PIs that supervised the project @itaytirosh.bsky.social, @suvalab.bsky.social, @roelverhaak.bsky.social, Antonio Iavarone and Anna Lasorella, all collaborators and institutions involved in the CARE consortium >
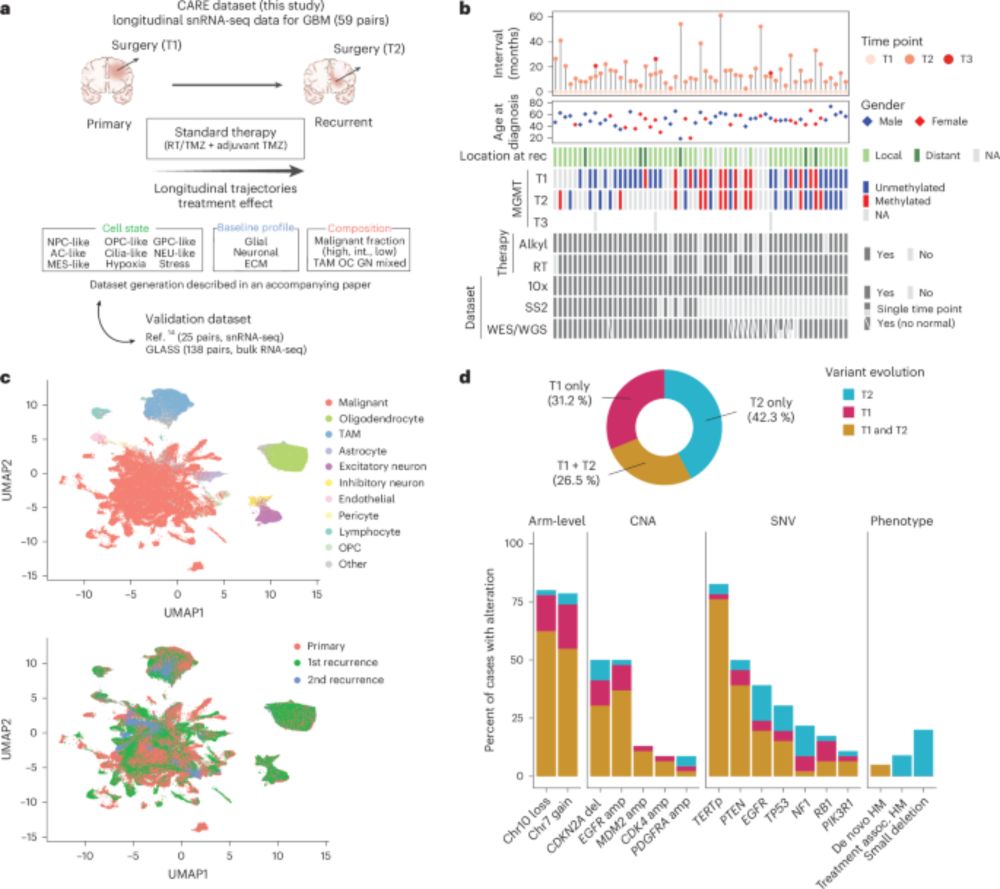
4/ And now for the results! Across the cohort the most consistent change at recurrence was reduced malignant cell fraction, with a reciprocal increase in glial and neuronal TME cells. This was observed in ~66% of patients, suggesting increased tumor integration into brain tissue.
3/ We first leveraged this large cohort to revisit the intra- and inter-tumor heterogeneity in GBM and characterized three multi-layered transcriptional ecosystems. Read more about this study in this great thread by @masashi-nomura.bsky.social →
bsky.app/profile/masa...
1/ Truly excited to share our new study that I had to privilege to co-lead during my PhD alongside great friends and collaborators @masashi-nomura.bsky.social, @kevin-johnson.bsky.social and Luciano Garofano, published at Nature Genetics @natureportfolio.nature.com!
www.nature.com/articles/s41...
I thank my great mentor/boss @suvalab.bsky.social and outstanding co-PIs @itaytirosh.bsky.social, @roelverhaak.bsky.social, Antonio Iavarone and Anna Lasorella. Also appreciate all collaborators, patients and their families who generously provided the samples.
12/ Before that, I would like to say thank wonderful co-first authors/friends, @avishayspitzer.bsky.social, @kevin-johnson.bsky.social and Luciano Garofano.
11/ How this architecture evolves during progression from primary to recurrent GBM is addressed in the second paper, that will be introduced by @avishayspitzer.bsky.social in the following threads later.

10/ These three layers of heterogeneity are inter-related and partially associated with specific genetic aberrations, thereby defining three stereotypic ecosystems in GBM. This work provides an unparalleled view of the multi-layered transcriptional architecture of GBM.

9/ Third, after controlling for the frequencies of cellular states, we find that the remaining variation between GBM samples highlights three baseline gene expression programs which we labeled Neuronal, Glial, and Extracellular Matrix.

8/ Second, we describe the diversity of cellular states and their pathway-based functional activities, with an expanded set of malignant cell states, including glial progenitor cell-like, neuronal-like, and cilia-like states that were previously depleted by tumor dissociation

7/ First, GBMs can be classified by their broad cellular composition, encompassing malignant, immune, vascular, neuronal, and glial cell types.
6/ In the first paper, we intensively analyzed GBM cellular heterogeneity, irrespective of time points. Our analyses reveled three transcriptomic layers that contribute to GBM heterogeneity.
5/ We divided the findings derived from this large-scale dataset into two papers.
4/ We thank @mdanderson.bsky.social, @dukecancer.bsky.social, @uoft.bsky.social, @pitiesalpetriere.bsky.social, NORLUX Neuro-Oncology laboratory, Seoul National University and @utokyoofficial.bsky.social for providing precious primary and recurrent GBM cohorts.
3/ In these studies, we collected 121 longitudinal GBM samples from 6 countries under GBM CARE (Cellular Analysis of Resistance and Evolution) consortium, and profiled them by single-nucleus RNA-seq (total 420 k cells).
2/These are the fantastic collaborative effort between Suva, Tirosh, Verhaak and Iavarone/Lasorella labs. We appreciate all supports from Mass General Brigham (Cancer Center, Pathology), @broadinstitute.org, Weizmann Institute of Science, @yalepress.bsky.social and University of Miami.

1/ Thrilled to share our TWO back-to-back papers published in Nature Genetics today! www.nature.com/articles/s41... , www.nature.com/articles/s41...