Congratulations Takashi!! 😃
Congratulations Takashi!! 😃
📣Huge congrats to our postdoc, Takaya Totsuka, on his new paper @currentbiology.bsky.social 🎉 He found that Ca2+ oscillations regulate oocyte spindle rotation! 🐭The project started during his PhD with Prof. Miho Ohsugi.
Free link: authors.elsevier.com/c/1lUqk3QW8S...

Having a lot of fun at #EMBOMobileGenome 😃
Perfect timing for our paper from the lab of @toddmacfarlan.bsky.social to be out @natcomms.nature.com!!
…and I’m currently on the job market looking for a new scientific home!
www.nature.com/articles/s41...

So excited to share this preprint from the Rosin lab in collaboration with the Hawley lab and @eelcotromer.bsky.social on moth spermatogenesis! We investigate the meiotic errors that occur during the formation of apyrene sperm (that have no DNA) in silkworms!
www.biorxiv.org/content/10.1...
🚨Our latest work on selfish centromeres is published @currentbiology.bsky.social 🐭🧬We found that the spindle checkpoint contributes to non-Mendelian segregation! Huge congrats to our lab manager, Zaak Walton, who led the project🎉
Free link:
authors.elsevier.com/a/1lQzN3QW8S...

Finally out! 🥳 Our paper showing how a transposable element (TE) insertion can cause developmental phenotypes is now published @natgenet.nature.com 🧬🦠🐁
Below is a brief description of the major findings. Check the full version of the paper for more details: www.nature.com/articles/s41588-025-02248-5
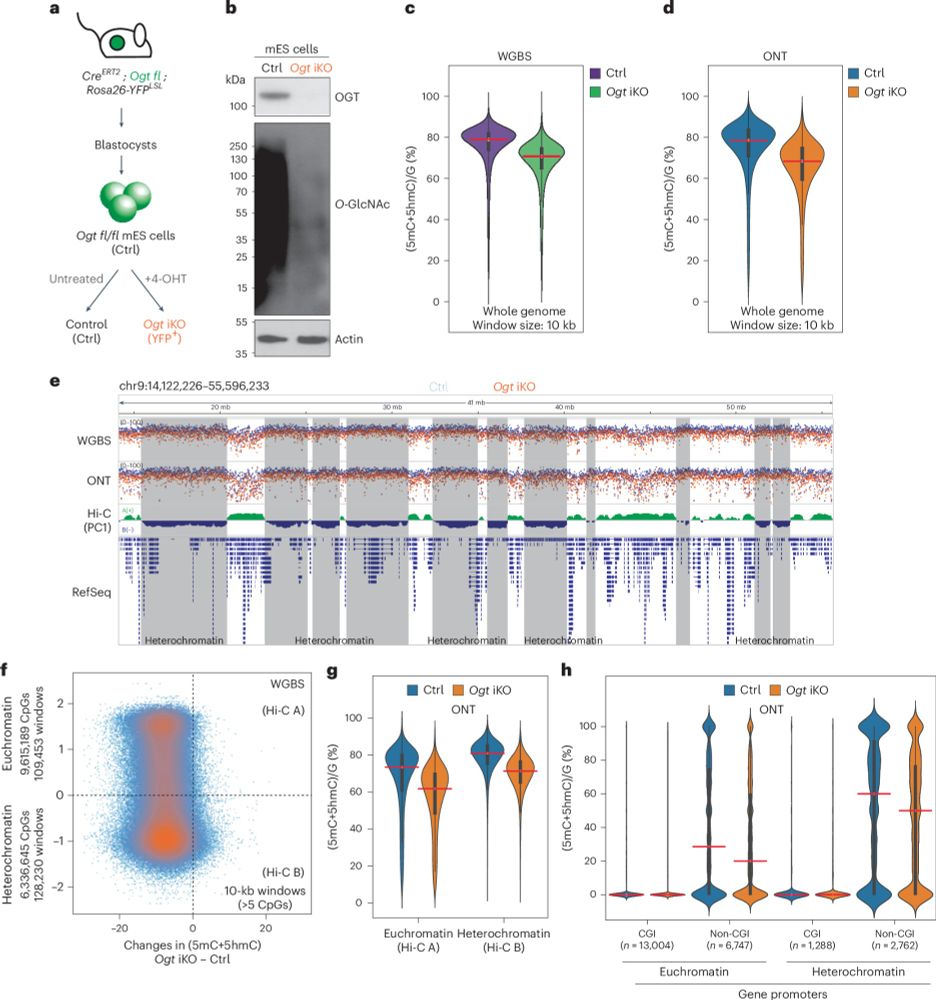
Our study with Anjana Rao's lab was just published at NSMB 🥳 Shows how OGT prevents TET proteins from activating retrotransposons, particularly those in heterochromatin. Paper has 6-base and ONT sequencing data from OGT iKO mESCs, and also interesting OGT inhibitor data 👀.
rdcu.be/efuho

New Akera's lab paper! Congrats @takashiakeralab.bsky.social and Eddie!
Huge thanks to all our collaborators for their help and for sharing expertise and data! This work would not have been possible otherwise!
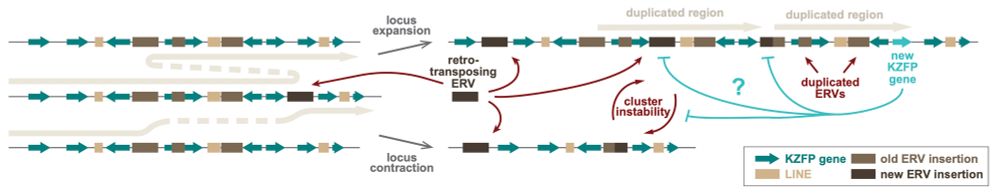
Our analyses also support a model by which new invading ERVs that landed at young KZFP gene clusters likely promoted the expansion and diversification of the KZFP gene repertoire in mice, by facilitating non-allelic homologous recombination events and large segmental duplications
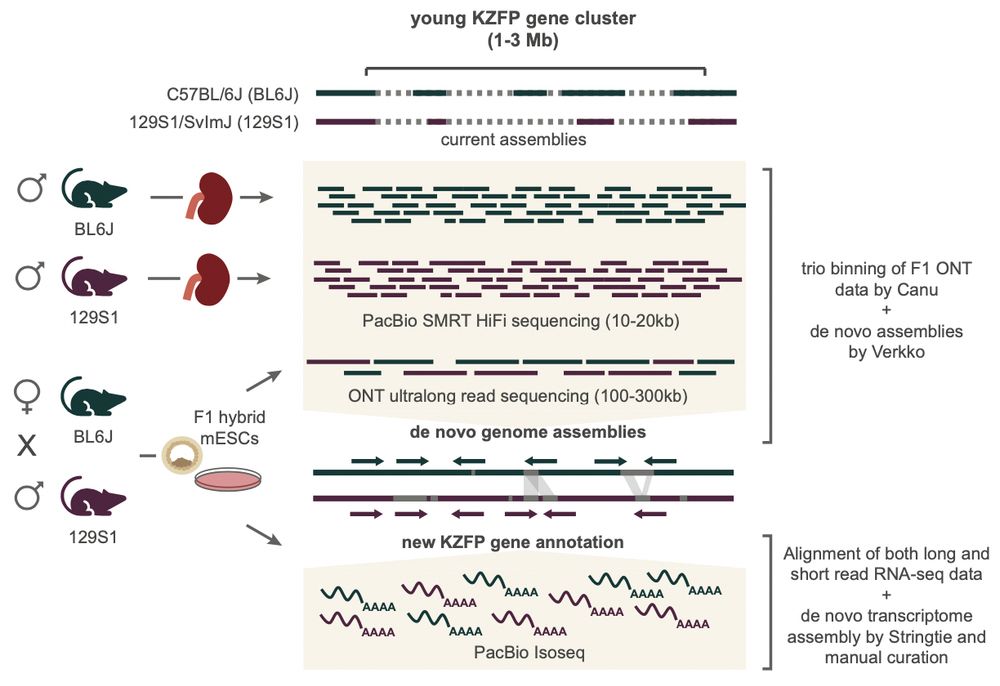
By combining multiple long-read sequencing technologies, we achieved a first detailed comparative analysis of young KZFP gene clusters in the mouse lineage, revealing a previously underestimated heterogeneity of these loci between mouse strains and species
I’m excited to share a new preprint from some of my work in the lab of @toddmacfarlan.bsky.social ! 😄
We dived into young KZFP gene clusters in different mouse strains and species to uncover mechanisms that facilitated their evolution and diversification in mice
Thank you Christa!!