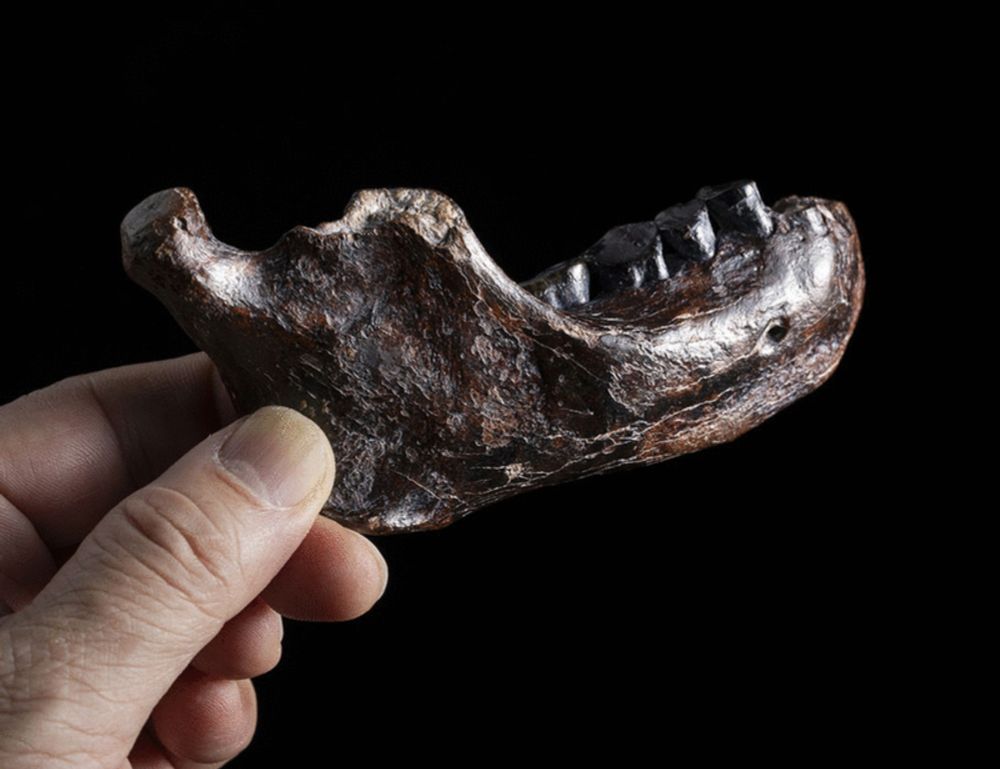
Given the recent and upcoming recovery of ancient proteins from archaic human remains this would be something to keep in mind. The missmatch of autosomal, mitochondrial, Y chromosome and morphology-based trees is already complicated for these groups, proteins might add some to that complexity!
02.03.2026 11:42
👍 1
🔁 0
💬 0
📌 0
While we suspect incomplete lineage sorting as the main culprit for this result, an older admixture between Neanderthal and humans, a 'super-archaic' admixture into Denisovans or other scenarios could be at play.
02.03.2026 11:42
👍 0
🔁 0
💬 1
📌 0
In fact, in the trees of enamel, collagen and other bone/dentin proteins, humans and Neanderthals appear closer to eachother as often as Neanderthals appear closer to Denisovans. This persists even when only looking at sub-saharan African populations, who have limited archaic introgression.
02.03.2026 11:42
👍 0
🔁 0
💬 1
📌 0
In short we find that while only a few proteins are needed to resolve the tree between humans - chimpanzees - gorillas, the same cannot be said for the tree of modern humans - Neanderthals - Denisovans.
02.03.2026 11:42
👍 0
🔁 0
💬 1
📌 0
🦷 A new study by @marinemorvan.bsky.social
High-throughput paleoproteomics method using MALDI-CASI-FTICR mass spectrometry to estimate biological sex from tooth enamel.
Great success on 130 individuals from medieval Great Moravia.
#Bioarchaeology #Archaeology #Proteomics #TooMS 😉 #Teammasspec
22.02.2026 12:59
👍 18
🔁 7
💬 0
📌 0
Seeing our work on videnskab.dk brings back memories of those fieldwork days. It was a great journey from sampling archaeological and old river sediments in Serbia to the ancient DNA labs in Copenhagen. Read more about the challenges and our findings in the original paper: doi.org/10.1038/s415...
23.02.2026 09:43
👍 4
🔁 1
💬 0
📌 0
Estén el pendiente y reserven la fecha 🤩🧬🇵🇪
20.02.2026 12:59
👍 1
🔁 3
💬 0
📌 0

we made a tool!
ybyra is a lightweight, flexible Y-chromosome haplogroup caller. It is faster, easier to tune and more robust than other tools particularly for low-coverage or ancient DNA data. And it has lots of detailed output – including plots!
www.biorxiv.org/content/10.1...
26.11.2025 08:29
👍 10
🔁 5
💬 1
📌 0
I'm hiring!⭐ As part of the DFF Sapere Aude-funded project MiddleEarth, I am looking for 1 postdoctoral researcher and 1 research assistant.
27.10.2025 09:54
👍 14
🔁 23
💬 1
📌 3
PAASTA Seminar Series - Ioannis Patramanis
YouTube video by PAASTA community
Following a marvelous time at ISBA11 and our PAASTA conference, we're back to sharing the work of our community with you here!
@ipatramanis.bsky.social PAASTA talk from March about PaleoProPhyler a tool for #phylogenies from ancient proteins is now live on our YouTube youtu.be/2gVEQyinL4A
02.09.2025 08:39
👍 14
🔁 7
💬 0
📌 0

Are you interested in doing a PhD in Copenhagen? Interested in studying Neanderthals and Denisovans which live on in our genomes?
Than you are more than welcome to apply to join my group starting Jan 2026 :)
candidate.hr-manager.net/ApplicationI...
Please reach out if you have any questions!
28.08.2025 12:35
👍 58
🔁 44
💬 1
📌 5

Questioning Credibility: Lynch’s Stem Cell Shortcomings
Vincent Lynch, a researcher at the University at Buffalo, has publicly raised doubts about the value and feasibility of de-extinction efforts.
The slander slanders on. Criticizing #ColossalBio and its co-founder Ben Lamm for their de-extinction #DisInformation campaign complete with #DeepFake dire wolves & woolly mammoths has unleashed a slew poorly written AI news stories about me, again 🙄
www.greenmatters.com/news/lynch-s...
07.07.2025 22:27
👍 10
🔁 2
💬 0
📌 0
The Paranthropus paper is out! :D
See the post from Palesa for a quick summary.
Although technically small in scale (4 specimens🦷), this work took many years and many people to complete...
Excited to see how the results we show will be applied to future sample sets, now that we know what works!
30.05.2025 08:50
👍 11
🔁 2
💬 1
📌 0

Inferring past demography and genetic adaptation in Spain using the GCAT cohort - Scientific Reports
Scientific Reports - Inferring past demography and genetic adaptation in Spain using the GCAT cohort
I am delighted to share with you some new insights into the selection processes that shaped the Spanish population!
This work has been led by @ebosch1972.bsky.social @francesccalafell.bsky.social ( @upf.edu ) in collaboration with Rafael de Cid ( @igtp.bsky.social ) and @sabiagini.bsky.social
⬇️⬇️⬇️
24.04.2025 15:38
👍 17
🔁 12
💬 1
📌 2
Nice! I wasn't aware of the paper, I'll be taking a look!
15.04.2025 20:58
👍 1
🔁 0
💬 1
📌 0
Well, thank you for reading it!
15.04.2025 13:03
👍 2
🔁 0
💬 0
📌 0
Lastly,
We comment on the current state of paleo-phylo-proteomics, including recent works, similar to ours (see academic.oup.com/gbe/article/... & www.researchsquare.com/article/rs-5...) and how the field can move forward.
15.04.2025 11:06
👍 1
🔁 0
💬 1
📌 0
The reason why trees differ, likely has to do with the amount of information lost, when going from mixed DNA data, to coding only DNA data (due to selection), to then protein data (due to translation). We quantify and visualize that informational loss.
15.04.2025 11:06
👍 0
🔁 0
💬 1
📌 0
DNA vs Proteins
Phylogenetic trees created using DNA or protein data can often differ from each other. But, does a tree generated from the protein and DNA data of the SAME LOCUS, differ? Turns out, yes, and in our small sample set, quite often (5 out of 12 genes).
15.04.2025 11:06
👍 1
🔁 0
💬 1
📌 0
What about more difficult relations?
We performed the same test as above for Neanderthal, Denisovans and modern humans. We failed to confidently or accurately resolve the relations between these 3 groups, with these 12 proteins. We added 16 additional proteins, but failed again to resolve them.
15.04.2025 11:06
👍 0
🔁 0
💬 1
📌 0
How many proteins do I need?
We see that around 3-4 proteins can give you enough variants to consistently discern between the 3 African great apes from one another (resolution) and around 9-10 proteins, accurately infer their phylogenetic relations, as we know them (accuracy).
15.04.2025 11:06
👍 0
🔁 0
💬 1
📌 0
How much phylogenetic information is in these proteins?
We saw that these proteins can vary greatly, with collagen and amelogenin (AMELX) being quite conserved, but other proteins (ODAM,COL17A1 and even AMELY!) being far more variable.
15.04.2025 11:06
👍 0
🔁 0
💬 1
📌 0
Our manuscript focuses on 12 enamel and collagen proteins that have consistently been recovered in samples older than 1 million years. We use extant and extinct hominids as our test model.
While this work still needs to go through peer review, here is a TLDR:
15.04.2025 11:06
👍 1
🔁 0
💬 1
📌 0


Hi
We contributed towards the South African Journal of Science Taung Centennial!
some links;
the special issue: issuu.com/sajs/docs/so...
our paper: sajs.co.za/article/view...
and I got featured on where I work from Nature Africa if you are interested in that: www.nature.com/articles/d44...
07.02.2025 10:46
👍 45
🔁 26
💬 2
📌 2

Paleoproteomics poster
Sequencing of ancient proteins is the only viable methodology today for retrieving the genetic data required to resolve evolutionary relations between vertebrate species that disappeared millions of years ago.
Paleoproteomics sheds light on million-year-old fossils 🏺🧪
www.nature.com/articles/s41...
11.12.2024 17:30
👍 33
🔁 4
💬 1
📌 0