
New preprint from
@novoalab.bsky.social !
Which tRNAs are used by ribosomes during translation?
We introduce tRIBO-seq, a nanopore method to sequence ribosome-associated tRNAs and track how the active tRNA pool changes across stress conditions.
www.biorxiv.org/content/10.6...
Thread 👇
04.03.2026 09:25
👍 35
🔁 13
💬 2
📌 1



Latest #CDlab paper on nanopore protein sequencing on @biorxivpreprint.bsky.social: www.biorxiv.org/cgi/content/...
Here, Justas Ritmejeris, @xiuqichen.bsky.social, collaborator @albadalab.bsky.social and me developed a conjugation chemistry strategy for nanopore sequencing of natural peptides!
13.11.2025 19:16
👍 32
🔁 8
💬 1
📌 1
Awesome place to do a PhD with exciting labs, a truly supportive PhD community, and the most scenic morning views you can imagine! 🌅
10/10
23.10.2025 09:34
👍 3
🔁 1
💬 0
📌 1
End-to-end protein design in the browser through evedesign. Generate and interactively explore designs in 2D/3D and export them as codon-optimized DNA. The underlying open source framework (released soon) is build to easily add new methods, more on that soon.
🌐 evedesign.bio
22.10.2025 14:30
👍 93
🔁 29
💬 2
📌 1
Very excited to share our new paper on the influence of biological sex and smoking in the clonal landscape of normal human bladder, just published in Nature.
A big thank you and congrats to all the authors, specially to @raquelbmi.bsky.social for all the shared efforts in this journey!
⬇️More info⬇️
08.10.2025 18:04
👍 18
🔁 6
💬 1
📌 0
The goal of a PhD is not to learn some facts or read a few papers or learn a bunch of techniques. The goal of a PhD is to learn independence, problem solving, how to finish things you start, resilience, & gain the ability to adapt & think creatively. Learning these things is hard.
13.08.2025 17:28
👍 337
🔁 87
💬 4
📌 5
Looks like private funders, HHMI,CZI etc., are not going to “save” US research. Europe is setting up mechanisms to attract US researchers & it may work. But, common, target junior talent, don’t shower money on superstars who will be flying professors & return to US when the science winter passes.
10.08.2025 15:35
👍 40
🔁 9
💬 0
📌 1


RNA N-glycosylation enables immune evasion and homeostatic efferocytosis by chemically caging acp3U. Excited to report this work lead by Vinnie @vinnieviruses.bsky.social and in collaboration with @vijayrathinam.bsky.social in @nature.com www.nature.com/articles/s41...
06.08.2025 15:36
👍 108
🔁 46
💬 7
📌 2
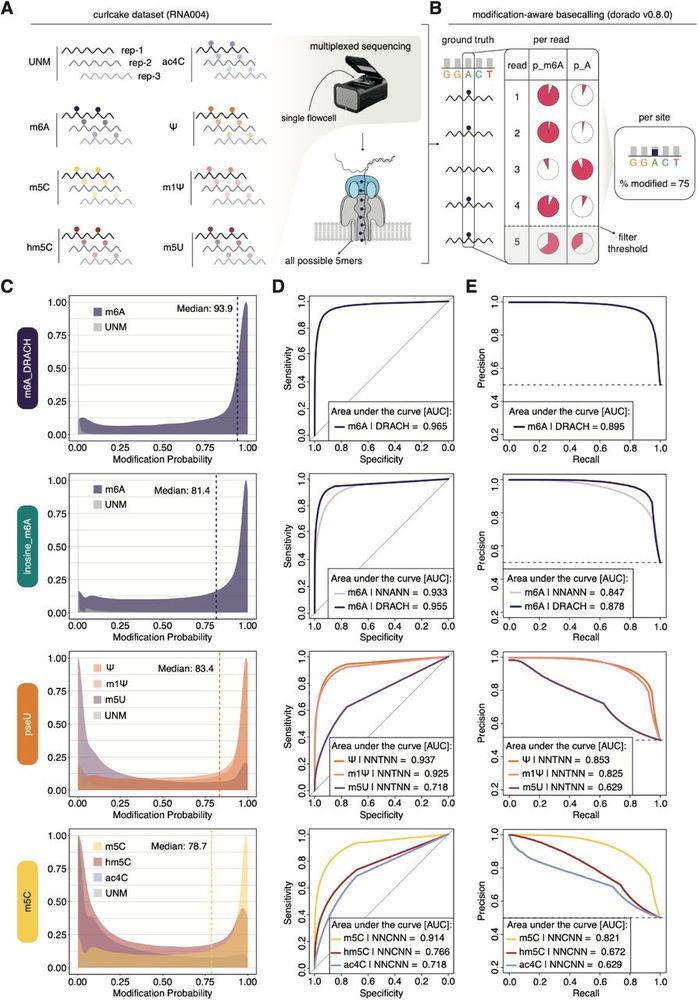
Systematic benchmarking of basecalling models for RNA modification detection with highly-multiplexed nanopore sequencing. #DirectRNAsequencing #DRS #RNAmodifications #Nanopore #Genomics #Bioinformatics #ModelBenchmarking #Basecalling @biorxivpreprint.bsky.social
www.biorxiv.org/content/10.1...
04.08.2025 19:05
👍 6
🔁 2
💬 0
📌 0
Moreover, for m1Y and Y, I am not aware of a non-enzymatic reaction that could convert one into the other during any downstream processing (leading to contamination). Unlike for m1A which can be converted to m6A at high temperatures, something that could happen easily at a heat inactivation step.
16.07.2025 11:28
👍 1
🔁 0
💬 0
📌 0
Hi James,
thanks for the nice words! I am speculating here but I think it is the latter. The training data should be synthetic sequences containing each of the modifications, and therefore quite clean.
16.07.2025 11:28
👍 1
🔁 0
💬 1
📌 0
Huge thanks to all the incredible people @novoalab.bsky.social that helped make this work possible! #rnasky #nanopore
14.07.2025 15:59
👍 4
🔁 0
💬 0
📌 0

Finally, we established that a combination of alignment and current features still presents a robust way of detecting RNA modifications with RNA004, as demonstrated on E.coli strains knockout for certain mod writers. (11/12)
14.07.2025 15:59
👍 3
🔁 0
💬 2
📌 0

While IVTs helped remove sequence-specific FPs two classes of FPs remained: I) cross-reactive mods found on the same base, and II) mods at adjacent positions (+/- 1nt) from identified FP sites, which cannot be accounted for with existing methods. (10/12)
14.07.2025 15:59
👍 1
🔁 1
💬 1
📌 0

Considering the substantial amount of FPs observed at unannotated sites we reasoned that the use of IVT controls might be necessary to help remove sequence specific FPs. (9/12)
14.07.2025 15:59
👍 1
🔁 0
💬 1
📌 0

From there we moved to rRNAs for benchmarking since accurate mod. maps exist for well-studied species. We could recapitulate most Ψ and m5C sites while also observing substantial FP, especially at base methylations and unannotated sites. (8/12)
14.07.2025 15:59
👍 1
🔁 0
💬 1
📌 0

Moving to per-site predictions (removing per read estimates that fall below the default cutoff), we observe that most models slightly underpredict the per-site stoichiometries with the pseU model performing the most accurate (particularly in the 0-25% modified range). (7/12)
14.07.2025 15:59
👍 1
🔁 0
💬 1
📌 0

Digging a bit deeper, we can identify particular sets of sequences (5mer) that produce lower probability estimates than the remaining ones, suggesting a strong impact of the sequence context on predictions. (6/12)
14.07.2025 15:59
👍 2
🔁 1
💬 1
📌 0

Key observations (per-read): 1) in general, models perform well. 2) Ψ and m1Ψ are indistinguishable. 3) m5C and hm5C cross-react in particular sequence contexts as indicated by the bimodal distribution of hm5C called with the m5C model. (5/12)
14.07.2025 15:59
👍 2
🔁 1
💬 2
📌 0

But first, a quick word on mod-aware basecallers and the information they provide. The models make predictions on a per-read and position basis (per-read) which can then be collapsed into per-site predictions, using downstream tools which consider a certain filter-threshold (per-site).
14.07.2025 15:59
👍 1
🔁 0
💬 1
📌 0

Next we went ahead and sequenced synthetic oligos containing all possible 5mers (n=1024) to assess model performance, cross-reactivities and potential sequence biases. (3/12)
14.07.2025 15:59
👍 2
🔁 0
💬 1
📌 0

First things first, we updated SeqTagger to support 96 barcodes with the newest sequencing chemistry (RNA004). If you are interested in multiplexing your own DRS runs the new model is openly available here -> github.com/novoalab/Seq.... Feedback is very welcome! (2/12)
14.07.2025 15:59
👍 2
🔁 0
💬 1
📌 0
Hätte mir vom Standard ein höheres Niveau erwartet, aber Klicks sind Klicks nehme ich an.
10.06.2025 17:54
👍 0
🔁 0
💬 0
📌 0

Endinsa’t en una vivència sensorial única que connecta ciència i art immersiu. “SEEING: Un ull i una caixa” arriba del 5 al 14 de juny al Centre Cívic Convent de Sant Agustí (Carrer del Comerç 36, Barcelona). www.crg.eu/en/event/see...
30.05.2025 11:24
👍 9
🔁 17
💬 0
📌 1
Very cool story and cover, congrats Eliana!
17.05.2025 17:57
👍 0
🔁 0
💬 0
📌 0




























