
The @steinaerts.bsky.social lab is looking for a postdoctoral researcher to develop next-generation sequence-to-function models for glioblastoma, one of the most aggressive brain cancers.
More info & how to apply 👉 https://vib.ai/en/opportunities#/job-description/130090
13.02.2026 10:02
👍 6
🔁 11
💬 0
📌 0
We are thrilled to share our new pre-print: “System-wide extraction of cis-regulatory rules from sequence-to-function models in human neural development”. S2F-deeplearning models can accurately encode enhancers, yet decoding these models into human-interpretable rules remains a major challenge.
15.01.2026 11:56
👍 44
🔁 21
💬 1
📌 1
Huge thanks to @steinaerts.bsky.social @s-poovathingal.bsky.social, Marta Wojno & @lukasmahieu.bsky.social for the support! @vibai.bsky.social & @vibneuroleuven.bsky.social
29.01.2026 14:31
👍 1
🔁 0
💬 0
📌 0
With more deep learning tools becoming available & easy to use to decode enhancer logic and cell-type identity, evaluating scATAC platforms of training data generation adds a new pillar to tech development! (10/10)
29.01.2026 14:25
👍 1
🔁 1
💬 0
📌 0
Technical consistency matters for integration:
We observe highly similar Tn5 insertion bias between HyDrop v2 and commercial droplet-based scATAC-seq platforms, across species. (9/n)
29.01.2026 14:24
👍 0
🔁 0
💬 1
📌 0

f, Computational design evaluating scATAC techniques as training data for S2F models in k-fold cross-validation (k=10). g-h, Scanning of ~2kb enhancers (g) with 500 bp sliding window (10bp
shift). Predicted accessibility of 500 bp windows of VT3067 enhancer (VDRC library). The region coordinates are based on Kvon et al. (2014), accessibility as ground truth. Line plots show mean predicted accessibility across n=10 cross-validation folds. Bands represent 95% confidence intervals (1.96 × SD). Both 10x and Hydrop v2-based models identify the same region as a core enhancer in the 200kb VDRC region. Taken from Figure 4.
f, S2F models in 10-fold cross-validation. g-h, Scanning of ~2kb enhancers with 500 bp sliding window. Predicted accessibility of neuroblast enhancer (VDRC library). Both 10x and Hydrop v2-based models identify the same region as a core enhancer in the 200kb VDRC region. Taken from Figure 4. (8/n)
29.01.2026 14:23
👍 0
🔁 0
💬 1
📌 0

d-e, 10x v1 and v2 datasets combined into the 10x Genomics training dataset compared to the HyDrop v2-based dataset for S2F deep learning
models in 10-fold cross-validation. Performance is validated on standard DL metrics and mouse cortex enhancers previously validated in vivo seen in e. (7/n)
29.01.2026 14:20
👍 0
🔁 0
💬 1
📌 0
Models trained on HyDrop v2 data accurately recover in vivo-validated enhancers in mouse and fly. Pretrained models are publicly available and ready for downstream use. (6/n)
29.01.2026 14:17
👍 0
🔁 0
💬 1
📌 0
We identify saturated sequencing depths for model training:
• ~12k reads/cell for Drosophila embryo
• ~36k reads/cell for mouse cortex
Beyond these points, deeper sequencing provides little added value. (5/n)
29.01.2026 14:16
👍 0
🔁 0
💬 1
📌 0
A central result:
Lower per-cell coverage can be offset by increasing dataset size, without loss of predictive performance. Even with ~60% more cells than 10x, HyDrop v2 datasets remain ~8× cheaper to generate. (4/n)
29.01.2026 14:16
👍 1
🔁 0
💬 1
📌 0

a, b, study design in mouse and Drosophila embryo. c, Estimation of generation costs of 67k cells in euros, excluding sequencing costs. HyDrop v1: 851.53 euro, HyDrop v2: 668.18 euro, 10x v2: 9,337.79
euro. Taken from Figure 1.
a, b, study design in mouse and Drosophila embryo. c, Estimation of generation costs of 67k cells in euros, excluding sequencing costs. HyDrop v1: 851.53 euro, HyDrop v2: 668.18 euro, 10x v2: 9,337.79
euro. Taken from Figure 1. (3/n)
29.01.2026 14:15
👍 1
🔁 0
💬 1
📌 0
With our new mouse motor cortex and Drosophila embryo data (600k cells!), we trained 220 sequence-to-function models, systematically varying read depth and cell number, and evaluated performance, robustness, and interpretability via cross-validation. (2/n)
29.01.2026 14:10
👍 0
🔁 0
💬 1
📌 0
We benchmarked custom vs commercial droplet-based scATAC-seq platforms for ML model training and introduce an optimized HyDrop v2 protocol with improved sensitivity and scalability. The full protocol is openly available. (1/n)
29.01.2026 14:09
👍 0
🔁 0
💬 1
📌 0
VIB-KU Leuven Center for Neuroscience
YouTube video by VIB-KU Leuven Center for Neuroscience
🚀 Proudly introducing the VIB-KU Leuven Center For Neuroscience, a merger of the two former VIB research centers VIB-KU Leuven Center for Brain & Disease Research and Neuro-Electronics Research Flanders (NERF)! Our new motto: Bold Science, Real Impact.
www.youtube.com/watch?v=uhaq...
27.01.2026 15:33
👍 15
🔁 7
💬 0
📌 1
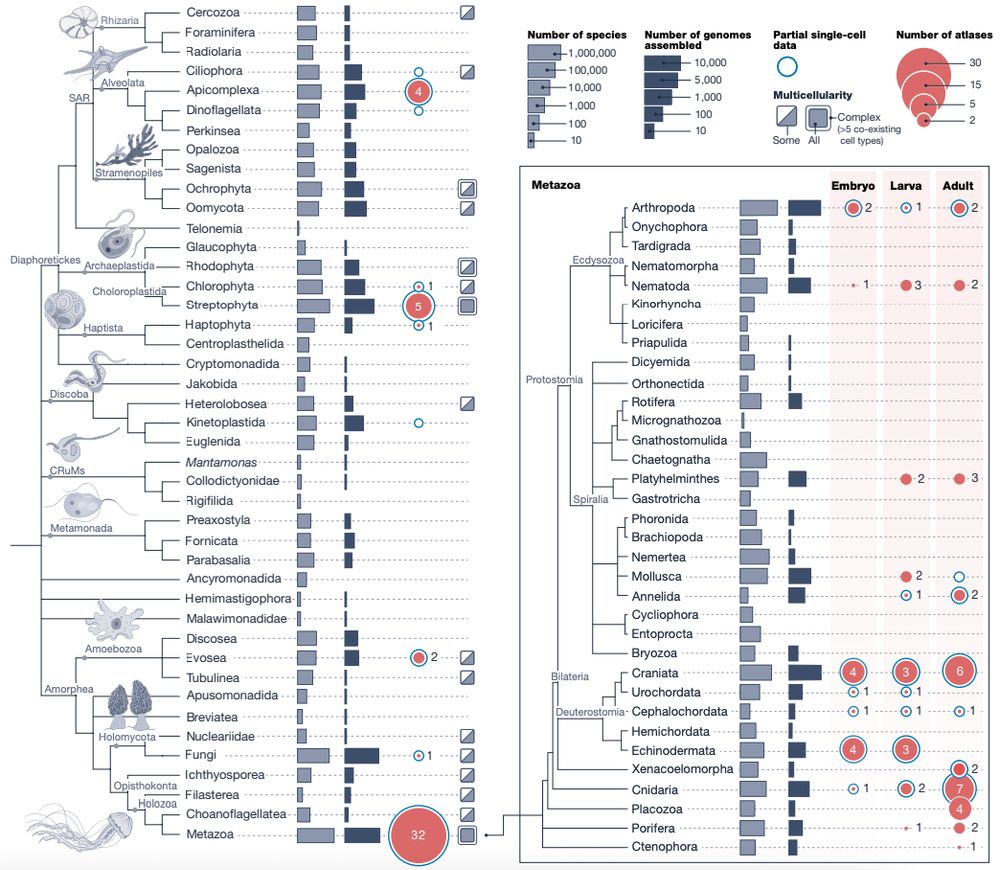
Happy to share the Biodiversity Cell Atlas white paper, out today in @nature.com. We look at the possibilities, challenges, and potential impacts of molecularly mapping cells across the tree of life.
www.nature.com/articles/s41...
24.09.2025 15:12
👍 228
🔁 106
💬 4
📌 10
Ending my two weeks conference marathon: last week Single Cell Spatial Omics #SCSO 600 year #KULeuven edition & now #SCG2025 - learned so much & am now already searching for the next one! Any recommendations for biodiversity, non-model organisms & genetics in 2026? 🪰🦋🐛🐌
18.09.2025 16:44
👍 4
🔁 0
💬 0
📌 0

ikea-style logo of splongget
1/ First preprint from @jdemeul.bsky.social lab 🥳! We present our new multi-modal single-cell long-read method SPLONGGET (Single-cell Profiling of LONG-read Genome, Epigenome, and Transcriptome)! www.biorxiv.org/content/10.1...
10.09.2025 15:48
👍 48
🔁 17
💬 1
📌 1
Make sure to also check out the preprint on the new CREsted package: many more sequence-to-function models across different species! … and it works with HyDrop v2 data as well.
Congrats to @niklaskemp.bsky.social & @seppedewinter.bsky.social & all the others working on this super cool project!
04.04.2025 10:04
👍 12
🔁 2
💬 0
📌 0
The data is out at GEO, resources.aertslab.org/papers/hydro..., and the models at crested.readthedocs.io/en/stable/mo... - we are looking forward to your feedback!
04.04.2025 09:58
👍 1
🔁 0
💬 0
📌 0
Thanks a lot to the whole lab for the great teamwork and especially to @s-poovathingal.bsky.social and @steinaerts.bsky.social for the support, Marta for the bead optimization, and @lukasmahieu.bsky.social for the help with all Deep Learning questions!
04.04.2025 09:05
👍 4
🔁 1
💬 1
📌 0
With HyDrop v2, we offer an accessible, scalable, and low-cost method to generate scATAC-seq datasets suitable for training ready-to-use deep learning tools. HyDrop v2 data reliably captures biologically relevant regulatory features and achieves predictive performance comparable to commercial tools.
04.04.2025 09:03
👍 2
🔁 0
💬 1
📌 0
Just as in fly, the mouse model trained on HyDrop v2 mouse cortex data also correctly identifies and matches predictions of in vivo validated enhancers.
04.04.2025 09:03
👍 0
🔁 0
💬 1
📌 0
We also compared our multi-class embryo model to the embryo models by Alexander Stark's lab @impvienna.bsky.social (sciATAC). With both 10x and HyDrop data-based sequence-to-function models, we were able to replicate and refine their motif predictions without the need for fine-tuning our models.
04.04.2025 09:01
👍 1
🔁 0
💬 1
📌 0
In the Drosophila embryo, we scored the in vivo validated enhancers from the VDRC lines and were able to track down the most predictive 500 bp region within the 2kb enhancers using our sequence-to-function models with 10-fold cross-validation.
04.04.2025 08:56
👍 0
🔁 0
💬 1
📌 0
We trained sequence-to-function models with the new CREsted package on both species and used the Seq2PRINT for footprinting in HyDrop and 10x data. Both are comparable as well in terms of enhancer predictions, sequence explainability, and transcription factor footprinting.
04.04.2025 08:55
👍 0
🔁 0
💬 1
📌 0
Differentially accessible regions & motif enrichment are equivalent between HyDrop v2 & 10x. Due to the bead optimization@s-poovathingal.bsky.social, HyDrop v2 experiments are very robust, and the data integrates seamlessly with commercially available chromatin accessibility methods (10x Genomics).
04.04.2025 08:54
👍 0
🔁 0
💬 1
📌 0
The costs of generating the library of, e.g., the mouse cortex across 35 experimental “runs” accumulate to merely 1,080.60 euros compared to an equivalent of 10x v2-based library preparation for 15,330.70 euros, excluding the sequencing cost. 35 experiments can be performed in just 1.5 days!
04.04.2025 08:53
👍 0
🔁 0
💬 1
📌 0








