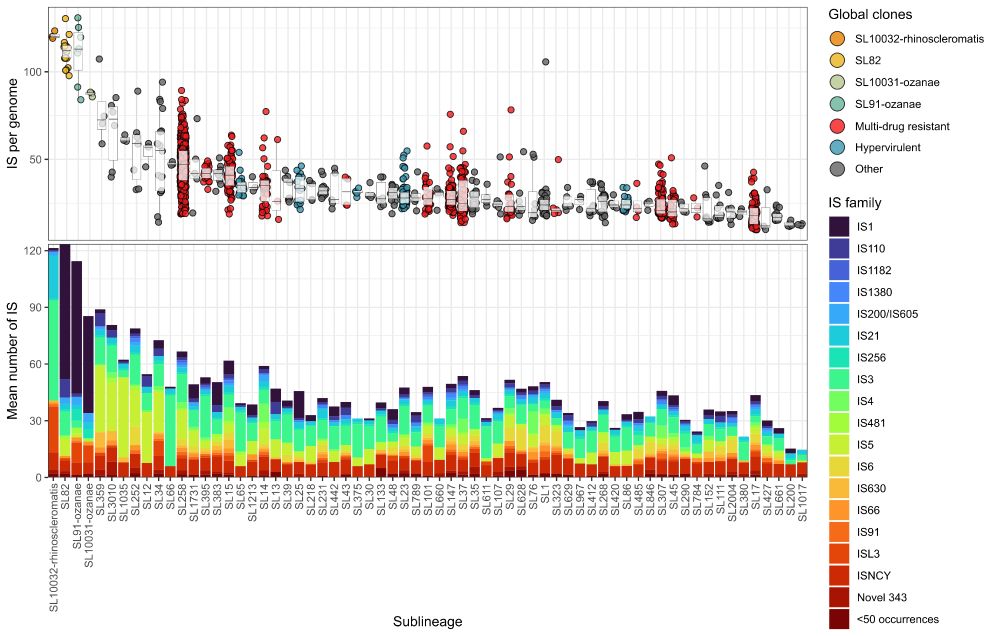
Graph showing IS loads by Klebsiella sub-lineage
New preprint where we analysed IS loads in 🦠 #Klebsiella pneumoniae lineages and found they're inversely associated with nutrient usage. We propose an insertional tolerance model, whereby IS can only insert into metabolically-tolerable sites.
#IDSky #MicroSky 🧪
www.biorxiv.org/content/10.1...
29.07.2025 02:29
👍 18
🔁 8
💬 1
📌 0
These findings show that Nanopore data is now accurate enough, practical and ready for further integration into clinical settings. Massive thankyou to @nenadmacesic.bsky.social, @yekwah.bsky.social, @katholt.bsky.social and all others who contributed to this project! /5
30.07.2025 02:14
👍 5
🔁 1
💬 0
📌 0
We provide specific recommendations for sequencing depth, basecalling model and analysis methods depending on the resources available and clinical context of the user. Analysis time is negligible, the biggest time savings come from choosing your basecalling model and depth requirements /4
30.07.2025 02:13
👍 6
🔁 0
💬 1
📌 0

We found modern ONT data (R10.4.1, Dorado SUP) to generate perfect MLST profiles and AMR allelic variant profiles, ~99.5% cgMLST profiles and cgSNP profiles highly concordant with traditional Illumina methods /3
30.07.2025 02:13
👍 4
🔁 0
💬 1
📌 0

Nanopore data has come a long way the last few years, and offers users a huge amount of flexibility in how they want to generate data. We took all combinations of sequencing chemistry, basecalling, depth & assembly status from 3 Gram-negative species groups and compared to hybrid gold standards /2
30.07.2025 02:11
👍 4
🔁 0
💬 1
📌 0

Pleased to say that our preprint benchmarking Nanopore data for MLST, cgMLST, cgSNP & AMR typing from bacterial isolates is out! TL;DR you can get almost perfect results from 50x depth using live SUP basecalling with a GPU in under 20 hours #microsky#IDsky 🦠🧬🖥️ /1
www.medrxiv.org/content/10.1...
30.07.2025 02:10
👍 46
🔁 29
💬 3
📌 2



