Lazarus lives!!
Lazarus lives!!
2 years of work packed into 2 little figures. When it finally worked, it 𝘸𝘰𝘳𝘬𝘦𝘥, beyond my wildest dreams. Try it out!

SOX2 and NR2F1 cooperate
to make #neurons in the #Thalamus
for visual input from the #retina
#DevBio
journals.biologists.com/bio/article/...
@biologyopen.bsky.social
In collab w Sara Mercurio and Silvia Nicolis @btbsunimib.bsky.social
Driven by Linda Serra and @annanordin.bsky.social
A 6-year long study from the Lab is finally out @genomeresearch.bsky.social
Mastered by @annanordin.bsky.social in collaboration with Chaitali Chakraborty from @remeseiro-lab.bsky.social
Why is our work Important? Discover it in the quick Bluetorial 👇
genome.cshlp.org/content/earl...
We had previously shown that TBX3 and #WNT cooperate in limb #DevBio
We now have a shocking finding, just published in @pnas.org
TBX3 physically engages with β-catenin to drive metastasis genes in #ColorectalCancer (CRC)
www.pnas.org/doi/epub/10....
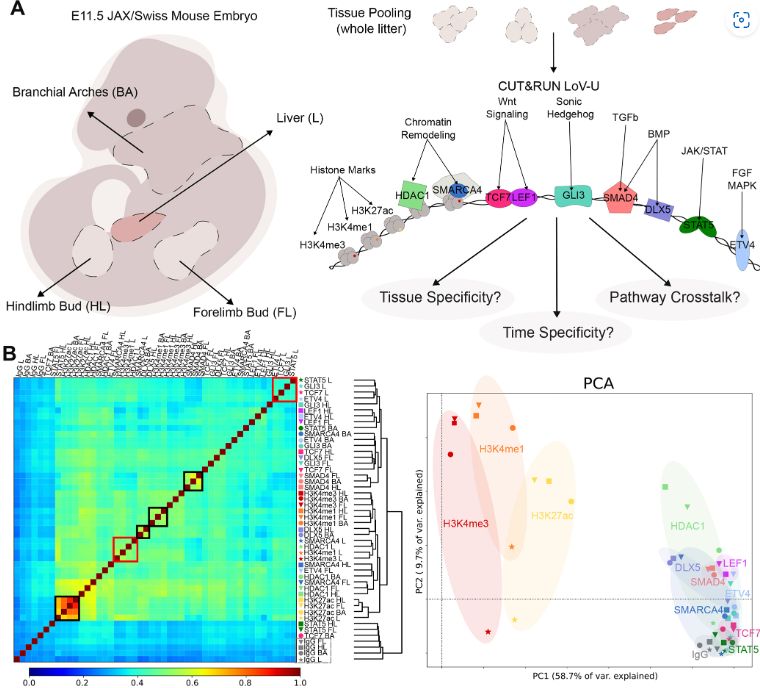
Figure 1 TFBomics of the E11.5 mouse embryo. (A) Schematic overview of the experimental design, detailing the tissues dissected from E11.5 JAX/Swiss mice and the targets profiled in each tissue using C&R LoV-U. (B) Left: Clustering of data according to Spearman's correlation. Black boxes highlight higher correlation or clustering according to the profiled factor, whereas red boxes highlight this between factors within the same tissue. IgG negative controls are highlighted in a gray box. Right: Overall data patterns shown by PCA. PC1 (58.7% of variance) separated some samples by the target factor (ellipses shaded by factor color code), while PC2 (9.7% of variance) typically separates the liver (marked with stars) from other tissues.
Construction of an atlas of transcription factor binding during mouse development identifies popular regulatory regions
Read this Techniques & Resources Article by @annanordin.bsky.social @gianlucazamba.bsky.social @claudiocantu81.bsky.social & co.: doi.org/10.1242/dev....
Our first release in the attempt to build the Transcription Factor Binding (TFB)-OMICS during #DevBio is now out in Development @dev-journal.bsky.social
doi.org/10.1242/dev....
👉We hope many will use our data, and some will want to help mapping their beloved gene regulators in their favorite tissue