A nice surprise for an otherwise troubling Monday morning, a volunteer profile describing my Peace Corps Jamaica service was in my inbox! Seeing the pictures of my students and plants in Jamaica made me smile. www.peacecorps.gov/what-we-do/l...
A nice surprise for an otherwise troubling Monday morning, a volunteer profile describing my Peace Corps Jamaica service was in my inbox! Seeing the pictures of my students and plants in Jamaica made me smile. www.peacecorps.gov/what-we-do/l...
🧪 🌱 It always puts me in a good mood when I see publications by colleagues and collaborators. Congrats Oussama and all the team for the work! MAGIC populations are a very powerful tool for genetics studies, this work will be very useful for the community! #SOLBreeding @upv.es 🍅🪄 #PlantScience

Time to buy a lottery ticket because I got today's Metaflora in a single guess! flora.metazooa.com?utm_source=m...
Actual footage of my plants trying to escape the growth chambers after I stress them. 🌱🌡️ #PlantScience #PlantStress
DOI link for the first article was dead for me but here it is if others are interested. www.biorxiv.org/content/10.6...
At the end of 2024 I did a chronological round up of all the #plantscience in @science.org that year.
So how did 2025 pan out? This year, I’m grouping papers thematically instead of chronologically so read on to find out what exciting plant science came out over the last 12 months. (1/22)
What a neat and informative figure! Looking forward to reading this review.

Phenotypic landscape of the velocity of the TS reaction (the rate-limiting step in DNA synthesis) as a function of activity of MTHFR and GNMT. A, wild-type conditions. Yellow sphere indicated wild-type; white spheres are mutations that lower the activity of the enzymes to 70 % and 35 % of wild-type. B, vitamin B12 deficiency (which lowers the activity of MS). A B12 deficiency destabilizes the TS reaction and the effects of mutations that increase TS activity are enhanced. (GNMT, glycine-N-methyltransferase; MTHFR, methylenetetrahydrofolate reductase; TS, thymidylate synthase).
#DBfeature
Cryptic genetic variation can be revealed by mutations or environmental factors that destabilize homeostatic mechanisms, producing phenotypic variants as a substrate for selection
By H. Frederik Nijhout
tinyurl.com/4dww8f3h
#SpecialIssue in research that transformed #DevBio

A pile of berry fruits including blueberries, strawberries, blackberries, raspberries, and goldenberries in the foreground.
New paper from friends pushing forward CRISPR improvement of an underutilized crop that I personally think is delicious, goldenberry! nph.onlinelibrary.wiley.com/doi/10.1002/... #plantscienceresearch
The 5th of 6 faculty positions in UNC Biology is posted. We're working with the @ncbg.bsky.social to fill a joint position as Director of the UNC Herbarium at @ncbg.bsky.social and as Associate Professor within the Department of Biology (BIOL). Please share 1/n
unc.peopleadmin.com/postings/308...
1/2 This manifesto is the result of a collaborative effort started earlier this year at the Plant Biology Education conference in Lancaster, UK.
We are delighted to see it out and we are proud that our Comms team contributed to such an important resource for the community! 🌱
I'd love some citations for for interest in TF change! I'm a cis-regulatory researcher dipping my toe into TFs, and it's much easier to find cis-regulatory papers (they tend to be lumped together, whereas TFs are called out by their specific names).

"With 2 weeks left before the end of this fiscal year, USDA’s [NIFA] had awarded just 558 competitive grants ... 68% fewer than during the prior fiscal year—and $741 million less in competitively awarded research funds"
Don't forget #PlantSci and ag research, folks.
www.science.org/content/arti...

A paper published in Nature Genetics presents a comprehensive pan-cistrome of maize leaves, revealing over 200,000 variants linked to transcription factor binding sites under both well-watered and drought conditions. go.nature.com/4785q5p 🧪🌾#plantscience
What a relevant justification! I can't wait to read the resulting issue!
Thanks for posting! I haven't seen your blog before and I really enjoyed the bountiful footnotes and citations!
Other authors, if you can find their contact information online. I know my inbox is less cluttered than senior folks and I try to respond promptly to requests!
An enjoyable read, with clear simple experiments that show tomato conservation of a core developmental mechanism previously demonstrated in Arabidopsis.
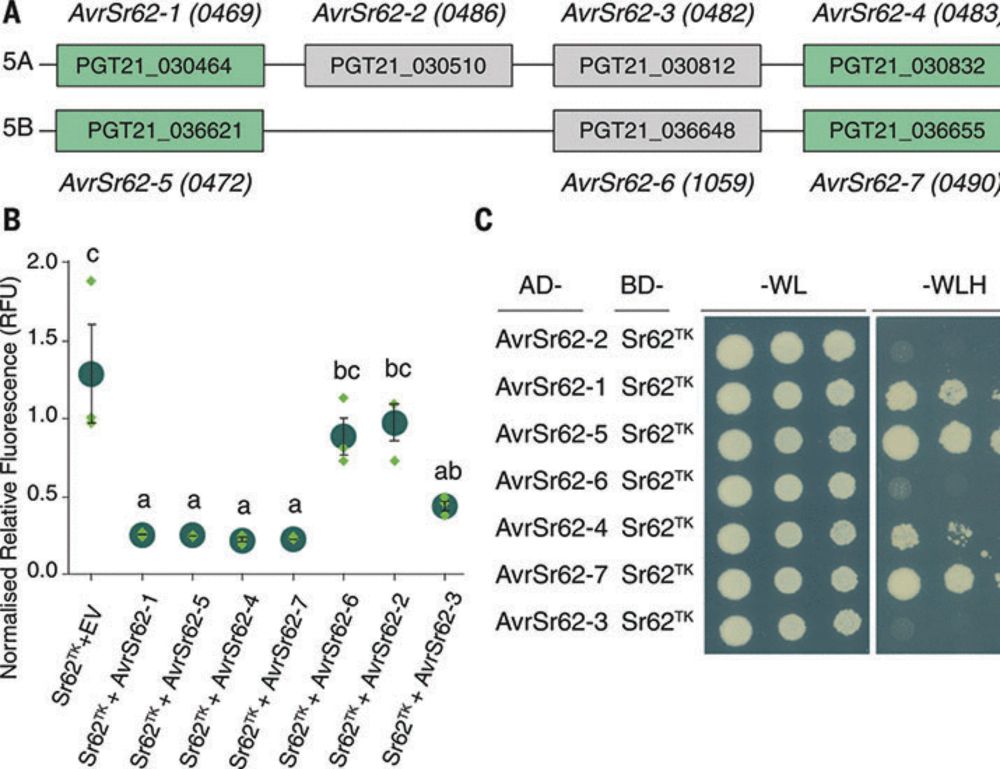
2 papers in @science.org this week uncover wheat tandem kinase function. Working in parallel, the authors of each paper show that two different tandem kinases converge to activate the same NLR!
www.science.org/doi/10.1126/...
www.science.org/doi/10.1126/...
#PlantScience
This paper is a tour de force from many friends and collaborators across Solanum expertise! Check it out to learn about the behavior of paralogs in genomes plus the Solanum genus and it's incredible potential for new crops! #plantscienceresearch
This team and our unique dataset have been instrumental in shaping my thinking about paralogs, epistasis, and quantitative genetics. I hope our work inspires others to reconsider the role of genomes in producing the diverse phenotypes of the natural world! #plantscienceresearch

Thrilled to share our latest preprint, an interdisciplinary look at epistasis between variants in the coding and noncoding genome! #plantscienceresearch www.biorxiv.org/content/10.1...
Nice work on evolution of genetic interaction between cis-reg elements in floral development! This study really hammered home for me the vast aesthetic difference between having one fewer petal (hideous) and one extra petal (charming) in brassicas. www.pnas.org/doi/10.1073/... #plantscienceresearch
Strong genetics and such a cool example of the consequences of altered TF-DNA binding in domestication!
Thanks for sharing! I enjoy the concept of "permitting yourself the feeling of doneness".

Diagram illustrating the 4 stages in DNA-free base editing: (1) in vitro transcription, (2) protoplast transfection, (3) protoplast regeneration, and (4) base-edited plants.
This remarkable new RNA-based adenine and cytosine #BaseEditing system achieves targeted and efficient A-to-G and C-to-T conversions in lettuce, with potential applications in #crop breeding and #PlantBiotechnology.
doi.org/10.1111/jipb...
#PlantScience
Associate Professor / Professor of Evolutionary Genomics
www.jobs.cam.ac.uk/job/49389/
Calling all #genomics researchers! The small, diverse, friendly Dept of Biological Sciences at Lehigh has a #tenuretrack position: bit.ly/3CYRb5B. App review starts 12/2 (not a hard cutoff). Please join our awesome team!